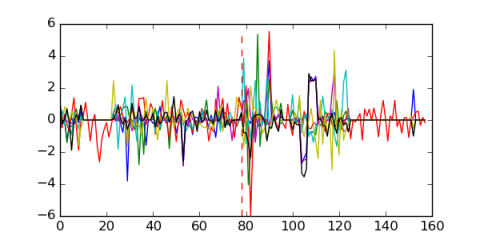
Module 574 Residual: 0.41
Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155)
| Residual | Motif 1 e-value | Motif 2 e-value | Conditions | Genes | version |
|---|---|---|---|---|---|
| 0.41 | 78 | 7 | 20160204 |
| Motif 1 | |
|---|---|
| Motif 2 | |
| MSMEG_1815 Uncharacterized protein |
KO_stationary.phase_rep1
KO_stationary.phase_rep2
Bedaquiline.vs..DMSO.15min
Mycobacterium.smegmatis.Spheroplast.induced
dosR.WT_Replicate4
Biological.replicate.4.of.7..ethambutol.treatment
Bedaquiline.vs..DMSO.30min
MB100pruR.vs..MB100..rep.3
Biological.replicate.5.of.6..negative.control
KO_log.phase_rep2
Biological.replicate.2.of.7..panaxydol.treatment
Plogcy5plogcy3slide110norm
DglnR_without.vs..with.N_rep1
MB100pruR.vs..MB100.rep.4
Biological.replicate.3.of.7..panaxydol.treatment
MB100pruR.vs..MB100.rep.2
stphcy5plogcy3slide487norm
WT.vs..DamtR.without.N_.dyeswap_rep2
MB100pruR.vs..MB100.rep.1
mc2155.vs.Dcrp1.replicate.3
Biological.replicate.1.of.6..kanamycin.treatment
rel.knockout.rep1
Bedaquiline.vs..DMSO..45min
DamtR.with.vs..DamtR.without.N_.rep1
JR121..pJR29.VapC..vs..JR121..pJR230..VapBC..rep3
Biological.replicate.4.of.6..kanamycin.treatment
DamtR.with.vs..DamtR.without.N_.rep2
X0.6..vs..50..air.saturation.Replicate.1
WT_log.phase_rep2
Sub.MIC.INH
X0.6..vs..50..air.saturation.Replicate.4
M.smegmatis_mc2_155_dMSMEG_0166_study_mutant_rep2
mc2155.vs.Dcrp1.replicate.2
Plogcy5plogcy3slide113norm
mc2155.vs.Dcrp1.replicate.1
mc2155.vs.Dcrp1.replicate.4
JR121..pJR29.VapC..vs..JR121..pJR230..VapBC..rep4
Biological.replicate.5.of.7..panaxydol.treatment
Stationary.cultures.control.without.induction.of.I.SceI.expression..T.0...Atc.
Plogcy5plogcy3slide111norm
stphcy5plogcy3slide580norm
X3dbfcy5plogcy3slide601norm
Stationary.phase.cultures.induced.to.express.I.SceI.for.15.minutes..T.15...ATc.
X3dbfcy3plogcy5slide607norm
mc2155.pMind..vs.mc2155.pMind.6189.replicate.4
X2.5..vs..50..air.saturation.Replicate.2
WT.with.vs..without.N_dye.swap_rep2
WT_stationary.phase_rep2
Log.phase.cultures.induced.to.express.I.SceI.for.45.minutes..T.45...ATc.
Biological.replicate.6.of.6..isoniazid.treatment
DamtR.with.vs..DamtR.without.N_dyeswap_.rep1
Biological.replicate.5.of.7..ethambutol.treatment
Wild.type.rep2.x
Slow.vs..Fast.Replicate.5
dosR.WT_Replicate3
Bedaquiline.vs..DMSO.60min
Biological.replicate.7.of.7..panaxydol.treatment
BatchCulture_LBTbroth_ExponentialPhase_Replicate3
X4dbfcy5plogcy3slide480norm
Wild.type.rep3
X4dbfcy5plogcy3slide901norm
mc2155.pMind..vs.mc2155.pMind.6189.replicate.1
Log.phase.cultures.without.induction.of.I.SceI.expression..T.0...Atc.
Biological.replicate.5.of.7..falcarinol.treatment
DglnR_without.vs..with.N_dyeswap_rep2
X2XR.rep1
DglnR_without.vs..with.N_dyeswap_rep1
JR121..pJR29.VapC..vs..JR121..pJR230..VapBC..rep1
BatchCulture_LBTbroth_StationaryPhase_Replicate4
JR121..pJR29.VapC..vs..JR121..pJR230..VapBC..rep2
BatchCulture_LBTbroth_StationaryPhase_Replicate2
BatchCulture_LBTbroth_StationaryPhase_Replicate3
BatchCulture_LBTbroth_StationaryPhase_Replicate1
Biological.replicate.2.of.6..kanamycin.treatment
M.smegmatis_mc2_155_dMSMEG_0166_study_wild.type_rep1
Biological.replicate.3.of.6..negative.control
X4dbfcy3plogcy5slide903norm
Bedaquiline.vs..DMSO.120min
Comments
Log in to post comments