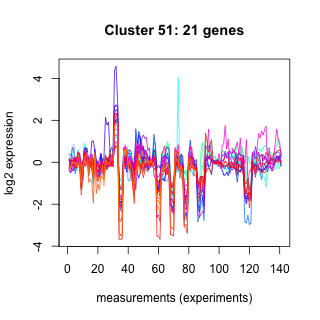
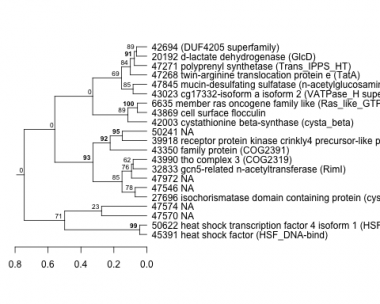
Phatr_hclust_0051 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0019343 | cysteine biosynthesis via cystathione | 0.00296369 | 1 | Phatr_hclust_0051 |
| GO:0006535 | cysteine biosynthesis from serine | 0.00886694 | 1 | Phatr_hclust_0051 |
| GO:0008299 | isoprenoid biosynthesis | 0.026385 | 1 | Phatr_hclust_0051 |
| GO:0008152 | metabolism | 0.051036 | 1 | Phatr_hclust_0051 |
| GO:0007264 | small GTPase mediated signal transduction | 0.0605726 | 1 | Phatr_hclust_0051 |
| GO:0015031 | protein transport | 0.0882177 | 1 | Phatr_hclust_0051 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.134605 | 1 | Phatr_hclust_0051 |
| GO:0006468 | protein amino acid phosphorylation | 0.406444 | 1 | Phatr_hclust_0051 |
| GO:0006118 | electron transport | 0.503223 | 1 | Phatr_hclust_0051 |
|
PHATRDRAFT_45391 : heat shock factor (HSF_DNA-bind) |
PHATRDRAFT_47845 : mucin-desulfating sulfatase (n-acetylglucosamine-6-sulfatase) (PRK13759) |
||
|
PHATRDRAFT_50622 : heat shock transcription factor 4 isoform 1 (HSF_DNA-bind) |
PHATRDRAFT_32833 : gcn5-related n-acetyltransferase (RimI) |
PHATRDRAFT_42003 : cystathionine beta-synthase (cysta_beta) |
PHATRDRAFT_47268 : twin-arginine translocation protein e (TatA) |
|
PHATRDRAFT_43990 : tho complex 3 (COG2319) |
PHATRDRAFT_43869 : cell surface flocculin |
PHATRDRAFT_47271 : polyprenyl synthetase (Trans_IPPS_HT) |
|
|
PHATRDRAFT_43350 : family protein (COG2391) |
PHATRDRAFT_6635 : member ras oncogene family like (Ras_like_GTPase superfamily) |
PHATRDRAFT_20192 : d-lactate dehydrogenase (GlcD) |
|
|
PHATRDRAFT_27696 : isochorismatase domain containing protein (cysteine_hydrolases superfamily) |
PHATRDRAFT_39918 : receptor protein kinase crinkly4 precursor-like protein (PKc_like superfamily) |
PHATRDRAFT_43023 : cg17332-isoform a isoform 2 (VATPase_H superfamily) |
PHATRDRAFT_42694 : (DUF4205 superfamily) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_0.5h | Re-illuminated_0.5h | -1.901 | 0.000396825 |
| BlueLight_0.5h | BlueLight_0.5h | -1.787 | 0.000462963 |
| GreenLight_0.5h | GreenLight_0.5h | -1.720 | 0.000458716 |
| Ammonia_SH | Ammonia_SH | -1.385 | 0.000344828 |
| RedLight_0.5h | RedLight_0.5h | -1.380 | 0.000555556 |
| Mixture_SH | Mixture_SH | -1.349 | 0.000357143 |
| highlight_12to48h | highlight_12to48h | -0.271 | 0.158159 |
| BlueLight_24h | BlueLight_24h | -0.210 | 0.146014 |
| GreenLight_24h | GreenLight_24h | -0.196 | 0.168788 |
| RedLight_6h | RedLight_6h | -0.170 | 0.470322 |
| RedLight_24h | RedLight_24h | -0.134 | 0.44 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.076 | 0.630205 |
| Green_vs_Red_24h | Green_vs_Red_24h | -0.063 | 0.618374 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | -0.059 | 0.638929 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.057 | 0.687376 |
| Re-illuminated_24h | Re-illuminated_24h | -0.025 | 0.886034 |
| Dispersed_oil_SH | Dispersed_oil_SH | -0.025 | 0.815096 |
| Si_free | Si_free | -0.016 | 0.91484 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.014 | 0.908042 |
| highlight_0to6h | highlight_0to6h | -0.006 | 0.981865 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 1 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | 0.040 | 0.720044 |
| Silver_SH | Silver_SH | 0.048 | 0.629625 |
| Oil_SH | Oil_SH | 0.056 | 0.590072 |
| Copper_SH | Copper_SH | 0.073 | 0.482203 |
| Dispersant_SH | Dispersant_SH | 0.079 | 0.464835 |
| Cadmium_0.12mg | Cadmium_0.12mg | 0.111 | 0.00875 |
| Cadmium_1.2mg | Cadmium_1.2mg | 0.114 | 0.118437 |
| light_16hr_dark_30min | light_16hr_dark_30min | 0.115 | 0.699194 |
| BlueLight_6h | BlueLight_6h | 0.130 | 0.461173 |
| light_15.5hr | light_15.5hr | 0.142 | 0.615135 |
| GreenLight_6h | GreenLight_6h | 0.189 | 0.295719 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | 0.200 | 0.347444 |
| Re-illuminated_6h | Re-illuminated_6h | 0.215 | 0.324104 |
| light_6hr | light_6hr | 0.254 | 0.285443 |
| Cadmium_SH | Cadmium_SH | 0.279 | 0.0320755 |
| Simazine_SH | Simazine_SH | 0.296 | 0.0226415 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | 0.301 | 0.133333 |
| Green_vs_Red_6h | Green_vs_Red_6h | 0.360 | 0.0598684 |
| light_10.5hr | light_10.5hr | 0.377 | 0.157819 |
| Dark_treated | Dark_treated | 1.866 | 0.000446429 |