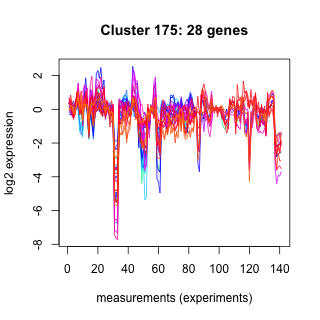
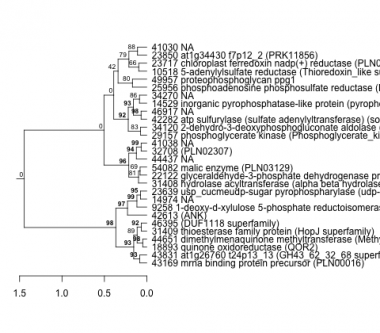
Phatr_hclust_0175 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006006 | glucose metabolism | 0.0239715 | 1 | Phatr_hclust_0175 |
| GO:0008299 | isoprenoid biosynthesis | 0.0307218 | 1 | Phatr_hclust_0175 |
| GO:0006306 | DNA methylation | 0.0670802 | 1 | Phatr_hclust_0175 |
| GO:0006725 | aromatic compound metabolism | 0.0670802 | 1 | Phatr_hclust_0175 |
| GO:0006118 | electron transport | 0.185759 | 1 | Phatr_hclust_0175 |
| GO:0005975 | carbohydrate metabolism | 0.197401 | 1 | Phatr_hclust_0175 |
| GO:0006096 | glycolysis | 0.00618972 | 1 | Phatr_hclust_0175 |
| GO:0000103 | sulfate assimilation | 0.00690418 | 1 | Phatr_hclust_0175 |
| GO:0008152 | metabolism | 0.0152246 | 1 | Phatr_hclust_0175 |
|
PHATRDRAFT_43169 : mrna binding protein precursor (PLN00016) |
PHATRDRAFT_9258 : 1-deoxy-d-xylulose 5-phosphate reductoisomerase (PRK05447) |
PHATRDRAFT_25956 : phosphoadenosine phosphosulfate reductase (PAPS_reductase) |
|
|
PHATRDRAFT_43831 : at1g26760 t24p13_13 (GH43_62_32_68 superfamily) |
PHATRDRAFT_23639 : usp_cucmeudp-sugar pyrophospharylase (udp-galactose glucose pyrophosphorylase) (PLN02830) |
PHATRDRAFT_29157 : phosphoglycerate kinase (Phosphoglycerate_kinase) |
PHATRDRAFT_49957 : proteophosphoglycan ppg1 |
|
PHATRDRAFT_18893 : quinone oxidoreductase (QOR2) |
PHATRDRAFT_31408 : hydrolase acyltransferase (alpha beta hydrolase superfamily) (Abhydrolase_6) |
PHATRDRAFT_34120 : 2-dehydro-3-deoxyphosphogluconate aldolase (KDPG_aldolase) |
PHATRDRAFT_10518 : 5-adenylylsulfate reductase (Thioredoxin_like superfamily) |
|
PHATRDRAFT_44651 : dimethylmenaquinone methyltransferase (Methyltransf_6 superfamily) |
PHATRDRAFT_22122 : glyceraldehyde-3-phosphate dehydrogenase precursor (GapA) |
PHATRDRAFT_42282 : atp sulfurylase (sulfate adenylyltransferase) (sopT) |
PHATRDRAFT_23717 : chloroplast ferredoxin nadp(+) reductase (PLN03116) |
|
PHATRDRAFT_31409 : thioesterase family protein (HopJ superfamily) |
PHATRDRAFT_54082 : malic enzyme (PLN03129) |
PHATRDRAFT_23850 : at1g34430 f7p12_2 (PRK11856) |
|
|
PHATRDRAFT_46395 : (DUF1118 superfamily) |
PHATRDRAFT_14529 : inorganic pyrophosphatase-like protein (pyrophosphatase) |
||
|
PHATRDRAFT_42613 : (ANK) |
PHATRDRAFT_32708 : (PLN02307) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_6h | Re-illuminated_6h | 0.041 | 0.877283 |
| Copper_SH | Copper_SH | -0.549 | 0.000515464 |
| Green_vs_Red_24h | Green_vs_Red_24h | 0.365 | 0.00441176 |
| Silver_SH | Silver_SH | 0.465 | 0.000555556 |
| Cadmium_1.2mg | Cadmium_1.2mg | -0.160 | 0.00915493 |
| BlueLight_24h | BlueLight_24h | -0.228 | 0.0581731 |
| GreenLight_0.5h | GreenLight_0.5h | -0.237 | 0.525075 |
| RedLight_6h | RedLight_6h | -0.299 | 0.123077 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 0.980594 |
| GreenLight_24h | GreenLight_24h | -0.100 | 0.526809 |
| light_6hr | light_6hr | 0.320 | 0.107831 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | 0.414 | 0.0189781 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.323 | 0.00128205 |
| Ammonia_SH | Ammonia_SH | -0.599 | 0.0158986 |
| Simazine_SH | Simazine_SH | -2.308 | 0.000423729 |
| Oil_SH | Oil_SH | -0.230 | 0.0276163 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | -0.573 | 0.000609756 |
| Cadmium_SH | Cadmium_SH | -0.088 | 0.402405 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | 0.237 | 0.104701 |
| highlight_0to6h | highlight_0to6h | 0.133 | 0.361787 |
| BlueLight_0.5h | BlueLight_0.5h | -0.527 | 0.211069 |
| Dispersant_SH | Dispersant_SH | 0.016 | 0.890929 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.555 | 0.000943396 |
| light_15.5hr | light_15.5hr | -1.768 | 0.000625 |
| light_10.5hr | light_10.5hr | -0.863 | 0.000561798 |
| Dispersed_oil_SH | Dispersed_oil_SH | 0.002 | 0.998586 |
| Si_free | Si_free | 0.124 | 0.353226 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | -0.250 | 0.578107 |
| Mixture_SH | Mixture_SH | -0.783 | 0.00231214 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | 0.301 | 0.080814 |
| light_16hr_dark_30min | light_16hr_dark_30min | -2.084 | 0.000735294 |
| BlueLight_6h | BlueLight_6h | 0.002 | 0.999499 |
| highlight_12to48h | highlight_12to48h | 0.353 | 0.03 |
| RedLight_24h | RedLight_24h | -0.465 | 0.00181818 |
| Dark_treated | Dark_treated | -4.316 | 0.000446429 |
| GreenLight_6h | GreenLight_6h | -0.321 | 0.0380952 |
| Cadmium_0.12mg | Cadmium_0.12mg | -0.010 | 0.854239 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.128 | 0.06625 |
| RedLight_0.5h | RedLight_0.5h | -1.836 | 0.000555556 |
| Green_vs_Red_6h | Green_vs_Red_6h | -0.022 | 0.928348 |
| Re-illuminated_24h | Re-illuminated_24h | -0.109 | 0.453406 |