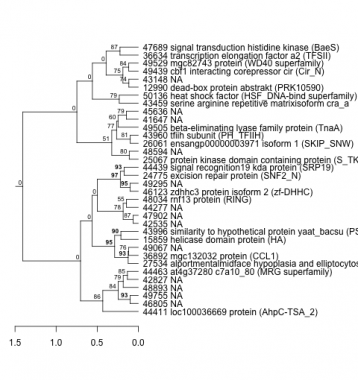
Phatr_hclust_0303 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006952 | defense response | 0.00370462 | 1 | Phatr_hclust_0303 |
| GO:0006354 | RNA elongation | 0.0110755 | 1 | Phatr_hclust_0303 |
| GO:0009072 | aromatic amino acid family metabolism | 0.0110755 | 1 | Phatr_hclust_0303 |
| GO:0006605 | protein targeting | 0.0220363 | 1 | Phatr_hclust_0303 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.0438421 | 1 | Phatr_hclust_0303 |
| GO:0016310 | phosphorylation | 0.0507136 | 1 | Phatr_hclust_0303 |
| GO:0006350 | transcription | 0.0751604 | 1 | Phatr_hclust_0303 |
| GO:0000160 | two-component signal transduction system (phosphorelay) | 0.0922612 | 1 | Phatr_hclust_0303 |
| GO:0000074 | regulation of cell cycle | 0.11901 | 1 | Phatr_hclust_0303 |
| GO:0006468 | protein amino acid phosphorylation | 0.131274 | 1 | Phatr_hclust_0303 |
|
PHATRDRAFT_44411 : loc100036669 protein (AhpC-TSA_2) |
PHATRDRAFT_15859 : helicase domain protein (HA) |
PHATRDRAFT_44439 : signal recognition19 kda protein (SRP19) |
PHATRDRAFT_43459 : serine arginine repetitive matrixisoform cra_a |
|
PHATRDRAFT_43996 : similarity to hypothetical protein yaat_bacsu (PSP1) |
PHATRDRAFT_25067 : protein kinase domain containing protein (S_TKc) |
PHATRDRAFT_50136 : heat shock factor (HSF_DNA-bind superfamily) |
|
|
PHATRDRAFT_12990 : dead-box protein abstrakt (PRK10590) |
|||
|
PHATRDRAFT_26061 : ensangp00000003971 isoform 1 (SKIP_SNW) |
|||
|
PHATRDRAFT_43960 : tfiih subunit (PH_TFIIH) |
PHATRDRAFT_49439 : cbf1 interacting corepressor cir (Cir_N) |
||
|
PHATRDRAFT_44463 : at4g37280 c7a10_80 (MRG superfamily) |
PHATRDRAFT_48034 : rnf13 protein (RING) |
PHATRDRAFT_49505 : beta-eliminating lyase family protein (TnaA) |
PHATRDRAFT_49529 : mgc82743 protein (WD40 superfamily) |
|
PHATRDRAFT_27534 : alportmentalmidface hypoplasia and elliptocytosis chromosomalgene 1 (AMMECR1) |
PHATRDRAFT_46123 : zdhhc3 protein isoform 2 (zf-DHHC) |
PHATRDRAFT_36634 : transcription elongation factor a2 (TFSII) |
|
|
PHATRDRAFT_36892 : mgc132032 protein (CCL1) |
PHATRDRAFT_47689 : signal transduction histidine kinase (BaeS) |
||
|
PHATRDRAFT_24775 : excision repair protein (SNF2_N) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_6h | Re-illuminated_6h | 0.018 | 0.946778 |
| Copper_SH | Copper_SH | -0.107 | 0.203079 |
| Green_vs_Red_24h | Green_vs_Red_24h | 0.185 | 0.0718085 |
| Silver_SH | Silver_SH | -0.109 | 0.168571 |
| Cadmium_1.2mg | Cadmium_1.2mg | 0.071 | 0.26 |
| BlueLight_24h | BlueLight_24h | -0.272 | 0.00892857 |
| GreenLight_0.5h | GreenLight_0.5h | 0.392 | 0.188211 |
| RedLight_6h | RedLight_6h | -0.648 | 0.000694444 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 1 |
| GreenLight_24h | GreenLight_24h | -0.240 | 0.0311828 |
| light_6hr | light_6hr | 0.211 | 0.272476 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | 0.276 | 0.0870283 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | -0.022 | 0.84225 |
| Ammonia_SH | Ammonia_SH | 0.693 | 0.00154321 |
| Simazine_SH | Simazine_SH | -0.024 | 0.801789 |
| Oil_SH | Oil_SH | -0.033 | 0.699208 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | 0.014 | 0.881328 |
| Cadmium_SH | Cadmium_SH | 0.089 | 0.363437 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | 0.153 | 0.224332 |
| highlight_0to6h | highlight_0to6h | 0.050 | 0.8385 |
| BlueLight_0.5h | BlueLight_0.5h | 0.995 | 0.00367647 |
| Dispersant_SH | Dispersant_SH | 0.111 | 0.213298 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.157 | 0.113348 |
| light_15.5hr | light_15.5hr | -0.187 | 0.37561 |
| light_10.5hr | light_10.5hr | 0.164 | 0.475668 |
| Dispersed_oil_SH | Dispersed_oil_SH | -0.021 | 0.825693 |
| Si_free | Si_free | 0.079 | 0.457971 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | 0.945 | 0.00296053 |
| Mixture_SH | Mixture_SH | 0.645 | 0.0051282 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | 0.430 | 0.0016129 |
| light_16hr_dark_30min | light_16hr_dark_30min | -0.256 | 0.19399 |
| BlueLight_6h | BlueLight_6h | -0.218 | 0.0940928 |
| highlight_12to48h | highlight_12to48h | -0.330 | 0.0267857 |
| RedLight_24h | RedLight_24h | -0.425 | 0.0025 |
| Dark_treated | Dark_treated | 1.388 | 0.00119048 |
| GreenLight_6h | GreenLight_6h | -0.197 | 0.160536 |
| Cadmium_0.12mg | Cadmium_0.12mg | -0.026 | 0.49461 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.032 | 0.701311 |
| RedLight_0.5h | RedLight_0.5h | -0.141 | 0.728141 |
| Green_vs_Red_6h | Green_vs_Red_6h | 0.452 | 0.0015625 |
| Re-illuminated_24h | Re-illuminated_24h | -0.218 | 0.0416201 |