Phatr_bicluster_0082 Residual: 0.37
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0082 | 0.37 | Phaeodactylum tricornutum |
Displaying 1 - 32 of 32
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_11438 undecaprenyl diphosphate synthase (Cis_IPPS)
PHATRDRAFT_12366 rna recognition motif familyexpressed (RRM_SF superfamily)
PHATRDRAFT_37341 acetolactate synthase (acolac_lg)
PHATRDRAFT_39062 (UBA)
PHATRDRAFT_41316 extracellular serine-threonine rich protein
PHATRDRAFT_42169 protein phosphatase (MPP_PP2A_PP4_PP6)
PHATRDRAFT_42871 extracellular threonine rich
PHATRDRAFT_43120 2-phosphoglycolate phosphatase (Gph)
PHATRDRAFT_44549 d-amino acid oxidase (DadA)
PHATRDRAFT_44835 mgc83623 protein (zf-C3HC4_3)
PHATRDRAFT_44838 mgc83623 protein (zf-C3HC4_3)
PHATRDRAFT_45770 mgc83491 protein (Thioredoxin_like superfamily)
PHATRDRAFT_45824 amino acid transporter (Aa_trans superfamily)
PHATRDRAFT_46648 platelet-binding glycoprotein
PHATRDRAFT_47710 subtilisin-like serine protease
PHATRDRAFT_48693 leucine-rich repeat family protein extensin family protein
PHATRDRAFT_49903 cystathionine gamma-synthase (MetC)
PHATRDRAFT_4994 calcium-dependent calmodulin-independent protein kinase cdpk-like (S_TKc)
PHATRDRAFT_51933 nonphototropic hypocotyl 1 (PAS)
PHATRDRAFT_8691 (6PGL)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006520 | amino acid metabolism | 0.0464863 | 1 | Phatr_bicluster_0082 |
| GO:0006865 | amino acid transport | 0.0540473 | 1 | Phatr_bicluster_0082 |
| GO:0008152 | metabolism | 0.0161272 | 1 | Phatr_bicluster_0082 |
| GO:0006468 | protein amino acid phosphorylation | 0.2936 | 1 | Phatr_bicluster_0082 |
| GO:0016567 | protein ubiquitination | 0.00814771 | 1 | Phatr_bicluster_0082 |

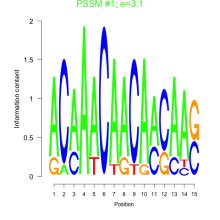
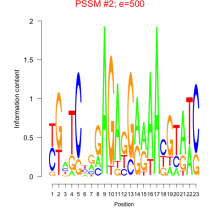
Comments