Phatr_bicluster_0116 Residual: 0.40
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0116 | 0.40 | Phaeodactylum tricornutum |
Displaying 1 - 27 of 27
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_16870 hmdh2_dicdi3-hydroxy-3-methylglutaryl-coenzyme a reductase 2 (hmg-reductase 2) (HMG-CoA_reductase_classI)
PHATRDRAFT_22166 zinc transporter (Zip superfamily)
PHATRDRAFT_33568 cytochrome p450 (p450 superfamily)
PHATRDRAFT_38294 (HSF)
PHATRDRAFT_39949 lpp3 (lipid phosphate phosphatase 3) (PAP2_like superfamily)
PHATRDRAFT_43594 pdz dhr glgf (DUF399 superfamily)
PHATRDRAFT_43878 ion transport protein (Ion_trans)
PHATRDRAFT_47930 histidine kinase response regulator hybrid protein (REC)
PHATRDRAFT_48179 myb-like dna-binding domain containing protein
PHATRDRAFT_48597 helicase domain protein (HA)
PHATRDRAFT_48747 (JmjC)
PHATRDRAFT_50059 cj022_human uncharacterized protein c10orf22 (DUF1637 superfamily)
PHATRDRAFT_50497 adenylate cyclase (CHD)
PHATRDRAFT_50607 methyltransferase protein (Methyltransf_21)
PHATRDRAFT_52551 n-acetylglucosaminyl transferase component (Gpi1 superfamily)
PHATRDRAFT_55209 (Carboxyl_trans)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006811 | ion transport | 0.0400529 | 1 | Phatr_bicluster_0116 |
| GO:0030001 | metal ion transport | 0.0577592 | 1 | Phatr_bicluster_0116 |
| GO:0009190 | cyclic nucleotide biosynthesis | 0.0612637 | 1 | Phatr_bicluster_0116 |
| GO:0006629 | lipid metabolism | 0.064756 | 1 | Phatr_bicluster_0116 |
| GO:0000160 | two-component signal transduction system (phosphorelay) | 0.0922612 | 1 | Phatr_bicluster_0116 |
| GO:0006812 | cation transport | 0.0956458 | 1 | Phatr_bicluster_0116 |
| GO:0000074 | regulation of cell cycle | 0.11901 | 1 | Phatr_bicluster_0116 |
| GO:0007242 | intracellular signaling cascade | 0.145022 | 1 | Phatr_bicluster_0116 |
| GO:0009058 | biosynthesis | 0.15459 | 1 | Phatr_bicluster_0116 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.19308 | 1 | Phatr_bicluster_0116 |

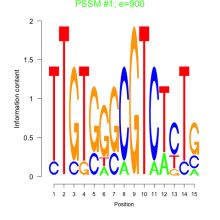
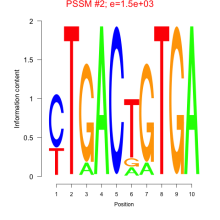
Comments