Phatr_bicluster_0192 Residual: 0.20
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0192 | 0.20 | Phaeodactylum tricornutum |
Displaying 1 - 32 of 32
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_11652 translation initiation factor if-3 (infC)
PHATRDRAFT_11653 nucleolar essential protein 1 (EMG1)
PHATRDRAFT_12140 chaperonedomain protein (DnaJ)
PHATRDRAFT_12896 ogd prolylhydroxylase domainfe ii 2- (Ofd1_CTDD)
PHATRDRAFT_12925 sumt_schpo probable uroporphyrinogen-iii c-methyltransferase (SUMT)
PHATRDRAFT_13281 mgc83122 protein (NIP7)
PHATRDRAFT_14176 transducin family protein wd-40 repeat family protein (WD40)
PHATRDRAFT_20773 loc733436 protein (CBF5)
PHATRDRAFT_2097 atp-dependent rna helicase ded1 (DEADc)
PHATRDRAFT_31987 density regulated protein drp1 (DENR_C)
PHATRDRAFT_33896 homeobox protein (PHD)
PHATRDRAFT_34956 g1 s-specific cyclin-e (COG5024)
PHATRDRAFT_34984 thyroid hormone receptor interactor 3
PHATRDRAFT_35395 cytochrome b5 (Cyt-b5)
PHATRDRAFT_37283 phosphoribosylglycinamide synthetase family protein (PRK07206)
PHATRDRAFT_42844 tiny fragments locus 9c (TrmA)
PHATRDRAFT_42926 programmed cell death 2-like (PDCD2_C superfamily)
PHATRDRAFT_44395 (RRM)
PHATRDRAFT_45828 sda1 domain containing 1 (SDA1)
PHATRDRAFT_45855 oxidoreductase-like protein (FabG)
PHATRDRAFT_46385 y2987_anasp upf0187 protein alr2987 (Bestrophin superfamily)
PHATRDRAFT_46837 nop14_mousenucleolar protein 14 (nucleolar complex protein 14) (Nop14)
PHATRDRAFT_47014 rna binding s1 domain protein (S1_like)
PHATRDRAFT_49586 phd-finger familyexpressed (RING)
PHATRDRAFT_4974 transducin wd-40 repeat protein (WD40 superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006915 | apoptosis | 0.00886694 | 1 | Phatr_bicluster_0192 |
| GO:0008152 | metabolism | 0.595805 | 1 | Phatr_bicluster_0192 |
| GO:0006779 | porphyrin biosynthesis | 0.0292769 | 1 | Phatr_bicluster_0192 |
| GO:0006457 | protein folding | 0.296964 | 1 | Phatr_bicluster_0192 |
| GO:0019538 | protein metabolism | 0.101759 | 1 | Phatr_bicluster_0192 |
| GO:0016567 | protein ubiquitination | 0.193944 | 1 | Phatr_bicluster_0192 |
| GO:0000074 | regulation of cell cycle | 0.0963646 | 1 | Phatr_bicluster_0192 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.487367 | 1 | Phatr_bicluster_0192 |
| GO:0007046 | ribosome biogenesis | 0.00886694 | 1 | Phatr_bicluster_0192 |
| GO:0006396 | RNA processing | 0.1541 | 1 | Phatr_bicluster_0192 |

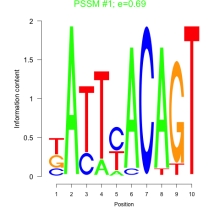
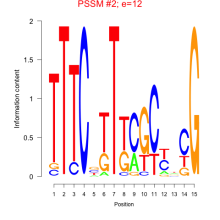
Comments