Regulatory Modules
Displaying 21 - 30 of 170Desulfovibrio vulgaris Hildenborough
| Title | Motif 1 | Motif 2 | Residual | Bicluster Genes | Selected GO Terms |
|---|---|---|---|---|---|
| bicluster_0191 |
0.0000000000000046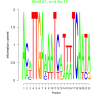 |
0.31 |
0.45 | DVU3122, DVU0273, DVU0303, DVU0304, DVU0762, DVU0763, DVU2377, DVU2378, DVU2379, DVU2380, DVU2383, DVU2388, DVU2389, DVU2390, DVU2571, DVU2572, DVU2573, DVU2574, DVU2680, DVU2681, DVU3331, DVU3332, DVU3333 | ion binding, metal ion binding, cation binding, transition metal ion binding, binding |
| bicluster_0219 |
0.12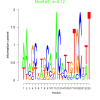 |
0.67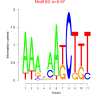 |
0.49 | DVU1082, DVU1678, DVU1964, DVU2066, DVU2067, DVU2265, DVU2648, DVU2651, DVU3146, DVU0695, DVU0781, DVU0782, DVU0783, DVU1393, DVU1859, DVU1965, DVU1966, DVU2017, DVU2652, DVU2653, DVU2917, DVU2919, DVU3138 | cellular protein modification process, protein modification process, macromolecule modification, 3'-5' exonuclease activity, hydrolase activity, acting on carbon-nit... |
| bicluster_0020 |
0.003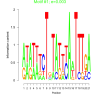 |
1 |
0.46 | DVU1542, DVU1543, DVU1565, DVU2624, DVU2646, DVU2647, DVU2702, DVU2703, DVU2719, DVU2720, DVU2721, DVU2722, DVU2723, DVU2724, DVU2727, DVU2728, DVU2729 | deaminase activity |
| bicluster_0343 |
0.0013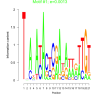 |
1.2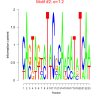 |
0.43 | DVU0259, DVU0267, DVU0056, DVU0074, DVU0222, DVU0268, DVU0281, DVU0945, DVU1190, DVU1470, DVU1762, DVU1796, DVU2227, DVU2443, DVU2847, DVU3039, DVU3182, DVU3375 | chemotaxis, taxis, locomotion, response to chemical, response to external stimulus |
| bicluster_0190 |
0.00000064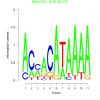 |
1.7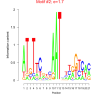 |
0.42 | DVU0371, DVU0554, DVU0555, DVU0556, DVU0558, DVU0559, DVU0560, DVU0561, DVU0562, DVU1147, DVU1152, DVU1166, DVU1167, DVU1947, DVU1948, DVU2010, DVU2175, DVU1692, DVU2174, DVU2177 | DNA integration, carbohydrate metabolic process, oxidoreductase activity, acting on the a... |
| bicluster_0272 |
0.12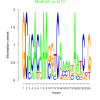 |
1.8 |
0.49 | DVU0129, DVU0222, DVU0690, DVU0720, DVU2443, DVU2633, DVU2799, DVU2800, DVU2801, DVU2847, DVU0629, DVU0677, DVU0722, DVU1440, DVU2180, DVU2351, DVU2747, DVU2835, DVU2933, DVU3016 | signal transducer activity, molecular transducer activity, phosphorelay sensor kinase activity, protein kinase activity, protein histidine kinase activity |
| bicluster_0243 |
0.17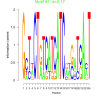 |
2.7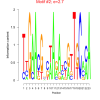 |
0.32 | DVU1707, DVU1059, DVU1063, DVU1700, DVU1733, DVU1747, DVU1996, DVU2122, DVU2186, DVU2207, DVU2262 | |
| bicluster_0148 |
0.029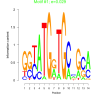 |
2.7 |
0.45 | DVU0174, DVU0220, DVU0253, DVU0278, DVU1569, DVU1816, DVU2202, DVU2557, DVU2968, DVU3263, DVU0006, DVU0138, DVU0238, DVU0305, DVU0883, DVU1568, DVU1812, DVU1904, DVU2076, DVU2583, DVU2616, DVU2684, DVU2935, DVU3227, DVU3296 | cellular homeostasis, homeostatic process, regulation of biological quality, biological regulation, copper ion binding |
| bicluster_0200 |
0.037 |
3 |
0.41 | DVU2457, DVUA0040, DVUA0045, DVUA0048, DVUA0055, DVUA0057, DVUA0058, DVUA0059, DVUA0060, DVUA0062, DVUA0063, DVU0086, DVU2458, DVU3105, DVUA0047, DVUA0141, DVUA0146 | polysaccharide biosynthetic process, polysaccharide metabolic process, cellular polysaccharide biosynthetic pro..., cellular carbohydrate biosynthetic proce..., cellular polysaccharide metabolic proces... |
| bicluster_0232 |
0.04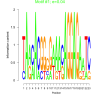 |
3.5 |
0.49 | DVU0302, DVU1190, DVU1372, DVU1373, DVU1374, DVU1375, DVU1377, DVU1378, DVU2532, DVU2965, DVU0837, DVU0838, DVU0936, DVU0942, DVU0986, DVU0987, DVU1093, DVU1367, DVU1392, DVU1612, DVU1834, DVU3187 | RNA processing, ncRNA processing, ncRNA metabolic process, cell cycle, TAT protein transport complex |

