Regulatory Modules
Displaying 101 - 110 of 170Desulfovibrio vulgaris Hildenborough
| Title | Motif 1 | Motif 2 | Residual | Bicluster Genes | Selected GO Terms |
|---|---|---|---|---|---|
| bicluster_0125 |
21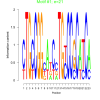 |
4.9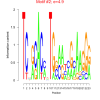 |
0.41 | DVU0310, DVU0311, DVU0312, DVU0313, DVU0314, DVU0315, DVU0512, DVU0513, DVU0515, DVU0516, DVU0517, DVU0519, DVU0913, DVU2232, DVU3003, DVU3018, DVU3019, DVU3020, DVU3004 | organelle organization, cell projection assembly, bacterial-type flagellum assembly, organelle assembly, cell projection organization |
| bicluster_0264 |
500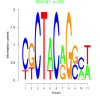 |
1300 |
0.40 | DVU1265, DVU2631, DVU2754, DVU0016, DVU0444, DVU1695, DVU2192, DVU2218, DVU2595, DVU2740, DVU2828, DVU3251, DVU3370 | [protein-PII] uridylyltransferase activi..., oxidoreductase activity, acting on NAD(P..., oxidoreductase activity, acting on NAD(P..., uridylyltransferase activity, nucleotidyltransferase activity |
| bicluster_0241 |
1.1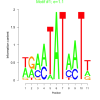 |
27 |
0.40 | DVU0170, DVU0174, DVU0729, DVU0820, DVU0821, DVU0822, DVU1263, DVU1774, DVU1811, DVU1813, DVU1814, DVU2304, DVU2345, DVU2585, DVU2615, DVU2676, DVU2833, DVU3155, DVUA0025 | aspartic-type endopeptidase activity, aspartic-type peptidase activity, endopeptidase activity, peptidase activity, acting on L-amino ac..., peptidase activity |
| bicluster_0163 |
1300 |
21000 |
0.40 | DVU0506, DVU1246, DVU1292, DVU1682, DVU1843, DVU2371, DVU3070, DVUA0006, DVUA0068, DVUA0076, DVU2051, DVU2372, DVU3088 | DNA recombination, DNA metabolic process, hydrolase activity, acting on carbon-nit..., hydrolase activity, acting on carbon-nit... |
| bicluster_0129 |
1000 |
2100 |
0.40 | DVU1496, DVU1498, DVU1499, DVU1500, DVU1502, DVU1514, DVU1522, DVU1524, DVU1493, DVU1513, DVU1515, DVU1516, DVU1517, DVU1525 | DNA (cytosine-5-)-methyltransferase acti..., DNA-methyltransferase activity, S-adenosylmethionine-dependent methyltra..., methyltransferase activity, transferase activity, transferring one-c... |
| bicluster_0003 |
480 |
2300 |
0.40 | DVU0230, DVU0426, DVU1482, DVU1699, DVU1957, DVU2158, DVU2249, DVU2296, DVU2343, DVU2432, DVU2567, DVU3078, DVU3249, DVU3251, DVUA0046 | cell wall organization, cell wall organization or biogenesis, external encapsulating structure organiz..., inorganic anion transmembrane transporte..., anion transmembrane transporter activity |
| bicluster_0295 |
1600 |
6800 |
0.40 | DVU0538, DVU1233, DVU3248, DVU0083, DVU0115, DVU1265, DVU1761, DVU2167, DVU2432, DVU2631, DVU2637, DVU2663, DVU2754, DVU2777, DVU3249 | transferase activity, transferring phosp..., [protein-PII] uridylyltransferase activi..., oxidoreductase activity, acting on NAD(P..., oxidoreductase activity, acting on NAD(P..., uridylyltransferase activity |
| bicluster_0036 |
28 |
1500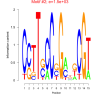 |
0.39 | DVU0081, DVU0083, DVU0316, DVU0484, DVU0494, DVU0514, DVU0636, DVU1433, DVU1940, DVU2663, DVU2865, DVU3034, DVU3106, DVU3387, DVU0486, DVU2622 | cofactor binding, ion binding, pyridoxal phosphate binding, FMN binding |
| bicluster_0171 |
120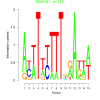 |
17000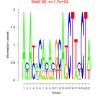 |
0.39 | DVU0698, DVU1075, DVU1301, DVU1427, DVU2200, DVU0817, DVU0886, DVU1778, DVU1849, DVU2179, DVU2215, DVU2220 | racemase and epimerase activity, racemase and epimerase activity, acting ..., endonuclease activity, cation transmembrane transporter activit... |
| bicluster_0038 |
47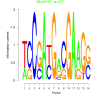 |
21000 |
0.39 | DVU0031, DVU2732, DVU2829, DVU2830, DVU2848, DVU2852, DVU2854, DVU2856, DVUA0014, DVUA0066, DVU0180, DVU0636, DVU2868 | enzyme regulator activity |

