|
bicluster_0343 |
0.0013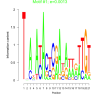 |
1.2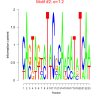 |
0.43 |
DVU0259, DVU0267, DVU0056, DVU0074, DVU0222, DVU0268, DVU0281, DVU0945, DVU1190, DVU1470, DVU1762, DVU1796, DVU2227, DVU2443, DVU2847, DVU3039, DVU3182, DVU3375 |
chemotaxis, taxis, locomotion, response to chemical, response to external stimulus |
|
mex_bicluster_0395 |
0.06 |
1.4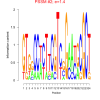 |
0.34 |
6830049, 6829520, 6825007, 6825733, 6830265, 6827818, 6829253, 6830941, 6825131, 6829794, 6826908, 6829519, 6827854, 6827143, 6827142, 6826990, 6830573, 6830999 |
|
|
mex_bicluster_0492 |
0.019 |
1.4 |
0.29 |
6825419, 6825337, 6828914, 6829790, 6827661, 6826476, 6831161, 6825456, 6828440, 6828096, 6830815, 6825875, 6828063, 6828062, 6829988, 6829874, 6827059, 6829874 |
|
|
mex_bicluster_0582 |
0.0000000000000059 |
1.5 |
0.34 |
6826381, 6827298, 6825595, 6828952, 6826202, 6827322, 6828967, 6828826, 6826462, 6828066, 6828947, 6827557, 6826307, 6826142, 6829591, 6829586, 6827103, 6831082, 6831037, 6830978, 6826485, 6830705, 6831183, 6831161, 6828807, 6826164, 6831147 |
|
|
mex_bicluster_0261 |
0.0000045 |
1.6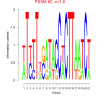 |
0.38 |
6830444, 6829311, 6830828, 6826505, 6830327, RMQ06697, 6826249, RMQ08303, 6829898, 6830640, RMQ09661, 6830227, 6828078, RMQ10914, 6831073, 6830331, 6828541, 6825053, 6825006, 6830736, 6827213, 6831141, 6828893, RMQ06580, 6829862, RMQ08028, RMQ08238, 6828905, 6830715, RMQ10222, 6830503, 6830911, 6825758 |
|
|
mmp-bicluster_0035 |
0.00069 |
1.6 |
0.43 |
MMP0077, MMP0219, MMP0383, MMP0590, MMP0591, MMP0592, MMP0593, MMP0718, MMP0719, MMP1073, MMP1190, MMP1191, MMP1493, MMP1518, MMP1519, MMP1520, MMP1560, MMP1561, MMP1562, MMP1563, MMP1564, MMP1565, MMP1566, MMP1567, MMP1643, MMP1644, MMP1717, MMP1718, MMP0076, MMP1373 |
cell wall, external encapsulating structure, transition metal ion binding, metal ion binding, cation binding |
|
bicluster_0190 |
0.00000064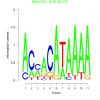 |
1.7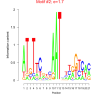 |
0.42 |
DVU0371, DVU0554, DVU0555, DVU0556, DVU0558, DVU0559, DVU0560, DVU0561, DVU0562, DVU1147, DVU1152, DVU1166, DVU1167, DVU1947, DVU1948, DVU2010, DVU2175, DVU1692, DVU2174, DVU2177 |
DNA integration, carbohydrate metabolic process, oxidoreductase activity, acting on the a... |
|
bicluster_0272 |
0.12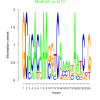 |
1.8 |
0.49 |
DVU0129, DVU0222, DVU0690, DVU0720, DVU2443, DVU2633, DVU2799, DVU2800, DVU2801, DVU2847, DVU0629, DVU0677, DVU0722, DVU1440, DVU2180, DVU2351, DVU2747, DVU2835, DVU2933, DVU3016 |
signal transducer activity, molecular transducer activity, phosphorelay sensor kinase activity, protein kinase activity, protein histidine kinase activity |
|
mex_bicluster_0238 |
0.000000066 |
1.9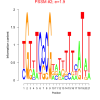 |
0.42 |
6829507, RMQ05638, 6825280, RMQ06015, 6827959, 6831200, 6829589, 6828442, 6828585, 6826470, 6827295, 6830873, 6829473, 6827671, 6826407, 6830300, 6826406, 6828374, 6825215, 6830358, RMQ11205, RMQ11300, 6827799 |
|
|
mmp-bicluster_0166 |
0.000017 |
2 |
0.30 |
MMP0018, MMP0097, MMP0238, MMP0538, MMP0547, MMP0558, MMP0652, MMP0756, MMP0758, MMP0759, MMP0813, MMP0842, MMP0864, MMP0871, MMP1268, MMP1291, MMP1292, MMP1294, MMP1296, MMP1604, MMP1605, MMP1653, MMP1654 |
pyruvate metabolic process, glycolytic process, carbohydrate catabolic process, single-organism carbohydrate catabolic p..., hydrolase activity, hydrolyzing O-glycos... |