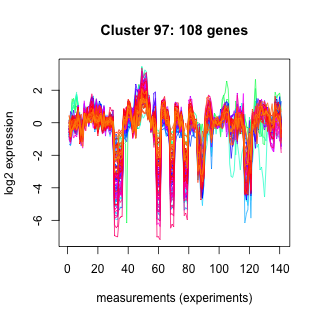
Phatr_hclust_0097 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0007067 | mitosis | 0.000100669 | 0.317308 | Phatr_hclust_0097 |
| GO:0051276 | chromosome organization and biogenesis | 0.000200608 | 0.629108 | Phatr_hclust_0097 |
| GO:0000074 | regulation of cell cycle | 0.000970899 | 1 | Phatr_hclust_0097 |
| GO:0006468 | protein amino acid phosphorylation | 0.00287233 | 1 | Phatr_hclust_0097 |
| GO:0030261 | chromosome condensation | 0.00592739 | 1 | Phatr_hclust_0097 |
| GO:0000278 | mitotic cell cycle | 0.00592739 | 1 | Phatr_hclust_0097 |
| GO:0007018 | microtubule-based movement | 0.013239 | 1 | Phatr_hclust_0097 |
| GO:0009186 | deoxyribonucleoside diphosphate metabolism | 0.0176813 | 1 | Phatr_hclust_0097 |
| GO:0016051 | carbohydrate biosynthesis | 0.0235082 | 1 | Phatr_hclust_0097 |
| GO:0006730 | one-carbon compound metabolism | 0.0235082 | 1 | Phatr_hclust_0097 |
|
PHATRDRAFT_47517 : gyltl1b protein (Glyco_transf_49 superfamily) |
PHATRDRAFT_44165 : (Smc) |
PHATRDRAFT_44270 : (SET) |
PHATRDRAFT_21410 : mitogen-activated protein kinase (STKc_MAPK) |
|
PHATRDRAFT_9158 : kinesin (kar3 subfamily) (Motor_domain superfamily) |
PHATRDRAFT_42743 : membrane protein (Rhomboid) |
||
|
PHATRDRAFT_49115 : (AlgLyase superfamily) |
PHATRDRAFT_50016 : lipopolysaccharide a protein (CAP10 superfamily) |
PHATRDRAFT_50280 : mgc81656 protein (DUF1032 superfamily) |
|
|
PHATRDRAFT_42993 : 200 kda antigen p200 (PTZ00121) |
PHATRDRAFT_32848 : mcm10_schpo dna replication licensing factor mcm10 (minichromosome maintenance protein 10) (cdc23 protein) |
||
|
PHATRDRAFT_49838 : myb-like dna-bindingshaqkyf class family protein (SANT) |
PHATRDRAFT_43956 : condensin subunit 1 (COG5098) |
||
|
PHATRDRAFT_15016 : transcription factor c-myb (SANT) |
PHATRDRAFT_47734 : proteophosphoglycan ppg4 |
PHATRDRAFT_48451 : (CHAT superfamily) |
|
|
PHATRDRAFT_43122 : protein regulator of cytokinesis 1 isoform 2 (MAP65_ASE1) |
PHATRDRAFT_8598 : b-type cyclin (CYCLIN) |
PHATRDRAFT_42914 : (PRK12323) |
|
|
PHATRDRAFT_38865 : cyclin d1 |
PHATRDRAFT_44947 : (PKc_like superfamily) |
PHATRDRAFT_46249 : (Alginate_lyase) |
|
|
PHATRDRAFT_38652 : (Motor_domain superfamily) |
PHATRDRAFT_2845 : protein serine threonine kinase (STKc_MAPKKK) |
PHATRDRAFT_13706 : long chain (FMN_red superfamily) |
|
|
PHATRDRAFT_47045 : viral a-type inclusion |
|||
|
PHATRDRAFT_50635 : (Alginate_lyase) |
PHATRDRAFT_49227 : (Anoctamin) |
PHATRDRAFT_42946 : (Got1) |
|
|
PHATRDRAFT_39117 : two component regulator three y domain protein |
PHATRDRAFT_13028 : sprt-like metalloprotease (SprT-like) |
PHATRDRAFT_44271 : (SET) |
|
|
PHATRDRAFT_47205 : (SANT) |
PHATRDRAFT_10628 : polo-like kinase 1 (S_TKc) |
PHATRDRAFT_48735 : chu large protein |
|
|
PHATRDRAFT_34026 : mkiaa0609 protein (Glyco_transf_49 superfamily) |
PHATRDRAFT_47992 : serine threonine protein kinase (PKc_like superfamily) |
||
|
PHATRDRAFT_47965 : wd repeat domain 18 (WD40 superfamily) |
PHATRDRAFT_36337 : myb-like dna-binding domain containing protein (SANT) |
PHATRDRAFT_38917 : dihydrofolate reductase (FSH1 superfamily) |
|
|
PHATRDRAFT_4344 : lrr protein |
|||
|
PHATRDRAFT_42574 : (alpha_CA superfamily) |
PHATRDRAFT_52174 : dna topoisomerase ii (PLN03128) |
||
|
PHATRDRAFT_50365 : myb-like dna-binding domain containing protein (SANT) |
|||
|
PHATRDRAFT_21455 : dynamin related protein involved in chloroplast division (Ras_like_GTPase superfamily) |
|||
|
PHATRDRAFT_43196 : metalloprotease mep1 homolog (ZnMc superfamily) |
PHATRDRAFT_36284 : loc398069 protein (Cnd2 superfamily) |
||
|
PHATRDRAFT_50429 : secreted lipopolysaccharide sugar transferase like family 8 glycosyltransferase (Glyco_tranf_GTA_type superfamily) |
PHATRDRAFT_47761 : glycoprotein 340 |
PHATRDRAFT_43757 : cyclin a |
|
|
PHATRDRAFT_47179 : excision repair cross-complementing rodent repair deficiency complementation group 6 - like (SNF2_N) |
|||
|
PHATRDRAFT_49507 : (Cnd1) |
PHATRDRAFT_23281 : nrs er (nucleotide-rhamnose synthase epimerase-reductase) (PLN02778) |
||
|
PHATRDRAFT_30352 : (Smc) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Ammonia_SH | Ammonia_SH | -2.342 | 0.000344828 |
| Mixture_SH | Mixture_SH | -2.596 | 0.000357143 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | -2.834 | 0.000396825 |
| Simazine_SH | Simazine_SH | 0.351 | 0.000423729 |
| Dark_treated | Dark_treated | -3.423 | 0.000446429 |
| GreenLight_0.5h | GreenLight_0.5h | -2.981 | 0.000458716 |
| BlueLight_0.5h | BlueLight_0.5h | -2.970 | 0.000462963 |
| Copper_SH | Copper_SH | 0.576 | 0.000515464 |
| highlight_12to48h | highlight_12to48h | 0.400 | 0.000555556 |
| RedLight_0.5h | RedLight_0.5h | -3.234 | 0.000555556 |
| light_10.5hr | light_10.5hr | 1.138 | 0.000561798 |
| light_6hr | light_6hr | 0.494 | 0.000561798 |
| BlueLight_6h | BlueLight_6h | 0.257 | 0.000581395 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | 0.556 | 0.000609756 |
| GreenLight_6h | GreenLight_6h | 0.247 | 0.000609756 |
| light_15.5hr | light_15.5hr | 1.808 | 0.000625 |
| Re-illuminated_24h | Re-illuminated_24h | 0.518 | 0.000657895 |
| Oil_SH | Oil_SH | 0.180 | 0.000666667 |
| RedLight_6h | RedLight_6h | 0.546 | 0.000694444 |
| light_16hr_dark_30min | light_16hr_dark_30min | 1.766 | 0.000735294 |
| Dispersant_SH | Dispersant_SH | -0.218 | 0.000909091 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | 0.935 | 0.000943396 |
| Green_vs_Red_6h | Green_vs_Red_6h | -0.299 | 0.0015625 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | -0.289 | 0.0016129 |
| Green_vs_Red_24h | Green_vs_Red_24h | -0.160 | 0.0075 |
| Si_free | Si_free | -0.173 | 0.00833333 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.163 | 0.0405405 |
| Silver_SH | Silver_SH | -0.102 | 0.0510822 |
| Cadmium_0.12mg | Cadmium_0.12mg | -0.033 | 0.065534 |
| Cadmium_SH | Cadmium_SH | 0.109 | 0.0710526 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | -0.155 | 0.0789604 |
| Dispersed_oil_SH | Dispersed_oil_SH | 0.073 | 0.211883 |
| highlight_0to6h | highlight_0to6h | 0.112 | 0.246948 |
| Cadmium_1.2mg | Cadmium_1.2mg | -0.040 | 0.252821 |
| GreenLight_24h | GreenLight_24h | -0.103 | 0.363203 |
| BlueLight_24h | BlueLight_24h | -0.106 | 0.466216 |
| RedLight_24h | RedLight_24h | 0.057 | 0.56662 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.010 | 0.879327 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.003 | 0.969433 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 0.980594 |
| Re-illuminated_6h | Re-illuminated_6h | -0.004 | 0.984909 |