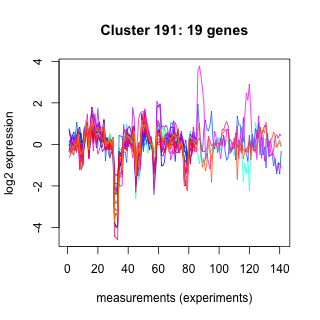
Phatr_hclust_0191 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0015988 | energy coupled proton transport, against electrochemical gradient | 0.00222277 | 1 | Phatr_hclust_0191 |
| GO:0019752 | carboxylic acid metabolism | 0.00444115 | 1 | Phatr_hclust_0191 |
| GO:0006754 | ATP biosynthesis | 0.00665514 | 1 | Phatr_hclust_0191 |
| GO:0009088 | threonine biosynthesis | 0.00665514 | 1 | Phatr_hclust_0191 |
| GO:0008033 | tRNA processing | 0.0436294 | 1 | Phatr_hclust_0191 |
| GO:0015986 | ATP synthesis coupled proton transport | 0.0793752 | 1 | Phatr_hclust_0191 |
| GO:0006396 | RNA processing | 0.117917 | 1 | Phatr_hclust_0191 |
| GO:0006457 | protein folding | 0.23215 | 1 | Phatr_hclust_0191 |
| GO:0008152 | metabolism | 0.492943 | 1 | Phatr_hclust_0191 |
|
PHATRDRAFT_19586 : spx (syg1 pho81 xpr1) domain-containing protein (SPX) |
PHATRDRAFT_14760 : alpha 2 frustulin |
PHATRDRAFT_35594 : thns2_ponpythreonine synthase-like 2 (Trp-synth-beta_II superfamily) |
PHATRDRAFT_43489 : helicase domain protein (HA) |
|
PHATRDRAFT_35363 : uba thif-type nad fad binding protein (COG1179) |
PHATRDRAFT_11965 : pseudouridinesynthase homolog 1 (PseudoU_synth superfamily) |
PHATRDRAFT_39209 : (SET) |
PHATRDRAFT_31662 : mono- or diacylglycerol acyltransferase (LPLAT superfamily) |
|
PHATRDRAFT_29608 : cg1109-isoform a (WD40) |
PHATRDRAFT_9706 : (COG2042) |
PHATRDRAFT_38362 : 2-isopropylmalate synthase (PRK12344) |
|
|
PHATRDRAFT_13386 : spermidine synthase (AdoMet_MTases) |
PHATRDRAFT_46062 : isoform cra_a |
PHATRDRAFT_24978 : h+lysosomal (vacuolar proton pump)beta 56 58isoform 2 (V-ATPase_V1_B) |
PHATRDRAFT_9070 : peptidyl-prolyl cis-transcyclophilin-type (cyclophilin superfamily) |
|
PHATRDRAFT_36899 : mitochondrial protein 18 kda (MTP18 superfamily) |
PHATRDRAFT_44310 : transmembrane protein (MS_channel superfamily) |
PHATRDRAFT_12921 : alm beta-like (S1_like superfamily) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Dark_treated | Dark_treated | -2.728 | 0.000446429 |
| light_6hr | light_6hr | -1.097 | 0.000561798 |
| BlueLight_0.5h | BlueLight_0.5h | -1.039 | 0.024 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | -0.972 | 0.00060241 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | -0.828 | 0.0560465 |
| light_10.5hr | light_10.5hr | -0.545 | 0.0505376 |
| Simazine_SH | Simazine_SH | -0.314 | 0.0222749 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | -0.247 | 0.0260274 |
| Cadmium_SH | Cadmium_SH | -0.169 | 0.17291 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | -0.148 | 0.542276 |
| Dispersant_SH | Dispersant_SH | -0.090 | 0.415607 |
| Silver_SH | Silver_SH | -0.083 | 0.394832 |
| Green_vs_Red_24h | Green_vs_Red_24h | -0.071 | 0.595192 |
| BlueLight_6h | BlueLight_6h | -0.065 | 0.766548 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.023 | 0.900223 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 1 |
| Cadmium_0.12mg | Cadmium_0.12mg | 0.005 | 0.969231 |
| Cadmium_1.2mg | Cadmium_1.2mg | 0.005 | 0.993249 |
| Dispersed_oil_SH | Dispersed_oil_SH | 0.006 | 0.974078 |
| Si_free | Si_free | 0.016 | 0.915718 |
| GreenLight_24h | GreenLight_24h | 0.042 | 0.8546 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | 0.048 | 0.623498 |
| Mixture_SH | Mixture_SH | 0.075 | 0.838925 |
| RedLight_6h | RedLight_6h | 0.083 | 0.755882 |
| BlueLight_24h | BlueLight_24h | 0.090 | 0.725497 |
| Green_vs_Red_6h | Green_vs_Red_6h | 0.100 | 0.711433 |
| RedLight_24h | RedLight_24h | 0.114 | 0.528829 |
| GreenLight_6h | GreenLight_6h | 0.183 | 0.346921 |
| highlight_0to6h | highlight_0to6h | 0.183 | 0.21125 |
| Re-illuminated_6h | Re-illuminated_6h | 0.207 | 0.377709 |
| Oil_SH | Oil_SH | 0.214 | 0.0680672 |
| Re-illuminated_24h | Re-illuminated_24h | 0.229 | 0.0984375 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | 0.237 | 0.0707819 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | 0.288 | 0.0386207 |
| GreenLight_0.5h | GreenLight_0.5h | 0.310 | 0.471955 |
| Copper_SH | Copper_SH | 0.348 | 0.00544872 |
| Ammonia_SH | Ammonia_SH | 0.570 | 0.0570144 |
| highlight_12to48h | highlight_12to48h | 0.584 | 0.00190476 |
| light_15.5hr | light_15.5hr | 0.658 | 0.0115385 |
| RedLight_0.5h | RedLight_0.5h | 0.690 | 0.0814433 |
| light_16hr_dark_30min | light_16hr_dark_30min | 0.716 | 0.00555556 |