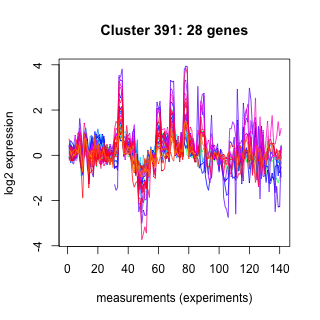
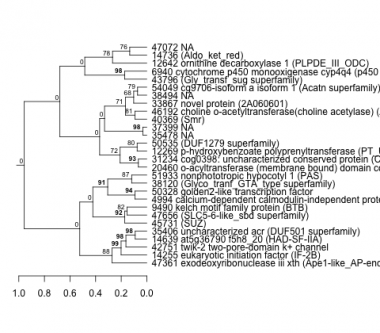
Phatr_hclust_0391 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006596 | polyamine biosynthesis | 0.00542673 | 1 | Phatr_hclust_0391 |
| GO:0006836 | neurotransmitter transport | 0.00542673 | 1 | Phatr_hclust_0391 |
| GO:0006298 | mismatch repair | 0.0162 | 1 | Phatr_hclust_0391 |
| GO:0006813 | potassium ion transport | 0.0530761 | 1 | Phatr_hclust_0391 |
| GO:0006413 | translational initiation | 0.0633789 | 1 | Phatr_hclust_0391 |
| GO:0006281 | DNA repair | 0.0886914 | 1 | Phatr_hclust_0391 |
| GO:0009058 | biosynthesis | 0.115816 | 1 | Phatr_hclust_0391 |
| GO:0006412 | protein biosynthesis | 0.360385 | 1 | Phatr_hclust_0391 |
| GO:0006468 | protein amino acid phosphorylation | 0.380037 | 1 | Phatr_hclust_0391 |
| GO:0006118 | electron transport | 0.473356 | 1 | Phatr_hclust_0391 |
|
PHATRDRAFT_47361 : exodeoxyribonuclease iii xth (Ape1-like_AP-endo) |
PHATRDRAFT_9490 : kelch motif family protein (BTB) |
PHATRDRAFT_12269 : p-hydroxybenzoate polyprenyltransferase (PT_UbiA_COQ2) |
|
|
PHATRDRAFT_14255 : eukaryotic initiation factor (IF-2B) |
PHATRDRAFT_4994 : calcium-dependent calmodulin-independent protein kinase cdpk-like (S_TKc) |
PHATRDRAFT_50535 : (DUF1279 superfamily) |
PHATRDRAFT_54049 : cg9706-isoform a isoform 1 (Acatn superfamily) |
|
PHATRDRAFT_42751 : twik-2 two-pore-domain k+ channel |
PHATRDRAFT_50328 : golden2-like transcription factor |
PHATRDRAFT_43796 : (Gly_transf_sug superfamily) |
|
|
PHATRDRAFT_14639 : at5g36790 f5h8_20 (HAD-SF-IIA) |
PHATRDRAFT_38120 : (Glyco_tranf_GTA_type superfamily) |
PHATRDRAFT_6940 : cytochrome p450 monooxigenase cyp4q4 (p450 superfamily) |
|
|
PHATRDRAFT_35406 : uncharacterized acr (DUF501 superfamily) |
PHATRDRAFT_51933 : nonphototropic hypocotyl 1 (PAS) |
PHATRDRAFT_40369 : (Smr) |
PHATRDRAFT_12642 : ornithine decarboxylase 1 (PLPDE_III_ODC) |
|
PHATRDRAFT_45731 : (SUZ) |
PHATRDRAFT_20460 : o-acyltransferase (membrane bound) domain containing 2 (PLN02332) |
PHATRDRAFT_46192 : choline o-acetyltransferase(choline acetylase) (ANK) |
PHATRDRAFT_14736 : (Aldo_ket_red) |
|
PHATRDRAFT_47656 : (SLC5-6-like_sbd superfamily) |
PHATRDRAFT_31234 : cog0398: uncharacterized conserved protein (COG0398) |
PHATRDRAFT_33867 : novel protein (2A060601) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_6h | Re-illuminated_6h | 0.430 | 0.0085034 |
| Copper_SH | Copper_SH | -0.541 | 0.000515464 |
| Green_vs_Red_24h | Green_vs_Red_24h | -0.089 | 0.39924 |
| Silver_SH | Silver_SH | 0.036 | 0.681869 |
| Cadmium_1.2mg | Cadmium_1.2mg | 0.005 | 0.983801 |
| BlueLight_24h | BlueLight_24h | 0.059 | 0.833598 |
| GreenLight_0.5h | GreenLight_0.5h | 1.430 | 0.000458716 |
| RedLight_6h | RedLight_6h | 0.412 | 0.0315625 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 0.980594 |
| GreenLight_24h | GreenLight_24h | 0.239 | 0.0475 |
| light_6hr | light_6hr | -0.425 | 0.0278689 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | -0.230 | 0.211174 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.069 | 0.483691 |
| Ammonia_SH | Ammonia_SH | 0.687 | 0.00752688 |
| Simazine_SH | Simazine_SH | -0.233 | 0.035041 |
| Oil_SH | Oil_SH | 0.049 | 0.594104 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | -0.138 | 0.183489 |
| Cadmium_SH | Cadmium_SH | -0.068 | 0.518009 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.269 | 0.0758065 |
| highlight_0to6h | highlight_0to6h | 0.176 | 0.16882 |
| BlueLight_0.5h | BlueLight_0.5h | 1.577 | 0.000462963 |
| Dispersant_SH | Dispersant_SH | -0.074 | 0.429802 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.492 | 0.000943396 |
| light_15.5hr | light_15.5hr | -1.246 | 0.000625 |
| light_10.5hr | light_10.5hr | -1.032 | 0.000561798 |
| Dispersed_oil_SH | Dispersed_oil_SH | 0.030 | 0.76488 |
| Si_free | Si_free | 0.137 | 0.315385 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | 1.763 | 0.000396825 |
| Mixture_SH | Mixture_SH | 0.545 | 0.0320565 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | -0.030 | 0.932109 |
| light_16hr_dark_30min | light_16hr_dark_30min | -1.040 | 0.000735294 |
| BlueLight_6h | BlueLight_6h | 0.382 | 0.00515873 |
| highlight_12to48h | highlight_12to48h | 0.178 | 0.343854 |
| RedLight_24h | RedLight_24h | 0.328 | 0.0186275 |
| Dark_treated | Dark_treated | 0.267 | 0.661757 |
| GreenLight_6h | GreenLight_6h | 0.313 | 0.0422535 |
| Cadmium_0.12mg | Cadmium_0.12mg | 0.021 | 0.636164 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.180 | 0.0111111 |
| RedLight_0.5h | RedLight_0.5h | 0.977 | 0.00280374 |
| Green_vs_Red_6h | Green_vs_Red_6h | -0.099 | 0.663308 |
| Re-illuminated_24h | Re-illuminated_24h | 0.295 | 0.00546219 |