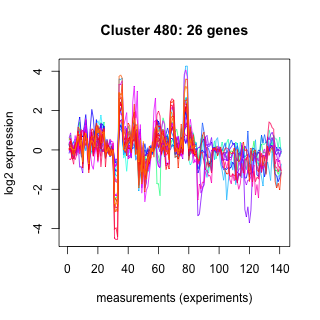
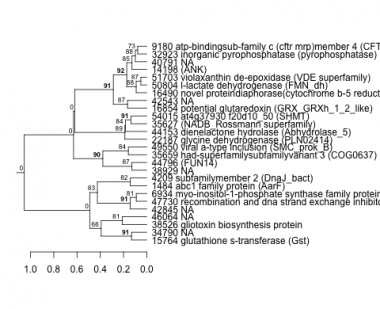
Phatr_hclust_0480 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006118 | electron transport | 0.00344047 | 1 | Phatr_hclust_0480 |
| GO:0006544 | glycine metabolism | 0.00886694 | 1 | Phatr_hclust_0480 |
| GO:0045454 | cell redox homeostasis | 0.0176618 | 1 | Phatr_hclust_0480 |
| GO:0006298 | mismatch repair | 0.0176618 | 1 | Phatr_hclust_0480 |
| GO:0006259 | DNA metabolism | 0.0350372 | 1 | Phatr_hclust_0480 |
| GO:0008152 | metabolism | 0.21493 | 1 | Phatr_hclust_0480 |
| GO:0006457 | protein folding | 0.296964 | 1 | Phatr_hclust_0480 |
| GO:0006810 | transport | 0.440472 | 1 | Phatr_hclust_0480 |
|
PHATRDRAFT_15764 : glutathione s-transferase (Gst) |
PHATRDRAFT_1484 : abc1 family protein (AarF) |
PHATRDRAFT_44153 : dienelactone hydrolase (Abhydrolase_5) |
PHATRDRAFT_50804 : l-lactate dehydrogenase (FMN_dh) |
|
PHATRDRAFT_4209 : subfamilymember 2 (DnaJ_bact) |
PHATRDRAFT_35627 : (NADB_Rossmann superfamily) |
PHATRDRAFT_51703 : violaxanthin de-epoxidase (VDE superfamily) |
|
|
PHATRDRAFT_38526 : gliotoxin biosynthesis protein |
PHATRDRAFT_54015 : at4g37930 f20d10_50 (SHMT) |
PHATRDRAFT_14198 : (ANK) |
|
|
PHATRDRAFT_44796 : (FUN14) |
PHATRDRAFT_16854 : potential glutaredoxin (GRX_GRXh_1_2_like) |
||
|
PHATRDRAFT_35659 : had-superfamilysubfamilyvariant 3 (COG0637) |
PHATRDRAFT_32923 : inorganic pyrophosphatase (pyrophosphatase) |
||
|
PHATRDRAFT_47730 : recombination and dna strand exchange inhibitor protein (ABC_ATPase superfamily) |
PHATRDRAFT_49550 : viral a-type inclusion (SMC_prok_B) |
PHATRDRAFT_16490 : novel proteindiaphorase(cytochrome b-5 reductase) (cyt_b5_reduct_like) |
PHATRDRAFT_9180 : atp-bindingsub-family c (cftr mrp)member 4 (CFTR_protein) |
|
PHATRDRAFT_6934 : myo-inositol-1-phosphate synthase family protein (NAD_binding_5 superfamily) |
PHATRDRAFT_22187 : glycine dehydrogenase (PLN02414) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_0.5h | Re-illuminated_0.5h | 2.095 | 0.000396825 |
| Simazine_SH | Simazine_SH | -0.649 | 0.000423729 |
| Dark_treated | Dark_treated | -1.880 | 0.000446429 |
| GreenLight_0.5h | GreenLight_0.5h | 1.252 | 0.000458716 |
| BlueLight_0.5h | BlueLight_0.5h | 2.040 | 0.000462963 |
| Copper_SH | Copper_SH | -0.473 | 0.000515464 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | -0.430 | 0.000609756 |
| Oil_SH | Oil_SH | -0.633 | 0.000666667 |
| Dispersed_oil_SH | Dispersed_oil_SH | -0.571 | 0.000675676 |
| Mixture_SH | Mixture_SH | -0.955 | 0.000675676 |
| light_16hr_dark_30min | light_16hr_dark_30min | -0.929 | 0.000735294 |
| Dispersant_SH | Dispersant_SH | -0.517 | 0.000909091 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.454 | 0.000943396 |
| highlight_12to48h | highlight_12to48h | 0.546 | 0.000980392 |
| Cadmium_SH | Cadmium_SH | -0.432 | 0.00120482 |
| light_6hr | light_6hr | 0.596 | 0.00147059 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | 0.565 | 0.00159574 |
| highlight_0to6h | highlight_0to6h | 0.312 | 0.00320513 |
| light_15.5hr | light_15.5hr | -0.643 | 0.00360825 |
| Re-illuminated_24h | Re-illuminated_24h | 0.331 | 0.00384615 |
| Ammonia_SH | Ammonia_SH | -0.619 | 0.0146512 |
| Silver_SH | Silver_SH | 0.231 | 0.0158046 |
| RedLight_6h | RedLight_6h | 0.451 | 0.0220149 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.194 | 0.0225352 |
| Green_vs_Red_6h | Green_vs_Red_6h | -0.365 | 0.0367187 |
| BlueLight_6h | BlueLight_6h | 0.280 | 0.06 |
| Re-illuminated_6h | Re-illuminated_6h | 0.309 | 0.0839662 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.123 | 0.0892473 |
| light_10.5hr | light_10.5hr | -0.371 | 0.121205 |
| RedLight_0.5h | RedLight_0.5h | 0.393 | 0.276575 |
| GreenLight_24h | GreenLight_24h | 0.145 | 0.322454 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | -0.171 | 0.373298 |
| RedLight_24h | RedLight_24h | 0.120 | 0.444498 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.099 | 0.477365 |
| Si_free | Si_free | 0.081 | 0.480702 |
| Cadmium_1.2mg | Cadmium_1.2mg | -0.042 | 0.615337 |
| GreenLight_6h | GreenLight_6h | 0.086 | 0.634422 |
| Green_vs_Red_24h | Green_vs_Red_24h | 0.025 | 0.867464 |
| Cadmium_0.12mg | Cadmium_0.12mg | -0.009 | 0.878133 |
| BlueLight_24h | BlueLight_24h | 0.022 | 0.954066 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 1 |