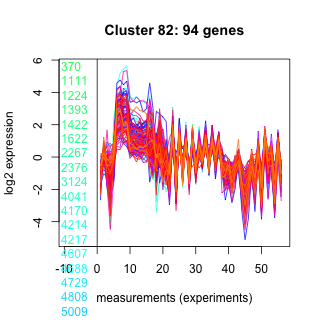
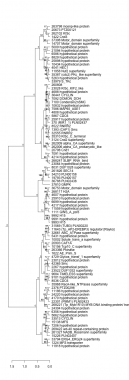
Thaps_hclust_0082 Hierarchical Clustering
Thalassiosira pseudonana
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0007018 | microtubule-based movement | 0.000000228 | 0.000835763 | Thaps_hclust_0082 |
| GO:0006334 | nucleosome assembly | 0.000000612 | 0.00223671 | Thaps_hclust_0082 |
| GO:0007001 | chromosome organization and biogenesis (sensu Eukaryota) | 0.0000105 | 0.0381485 | Thaps_hclust_0082 |
| GO:0016310 | phosphorylation | 0.00789406 | 1 | Thaps_hclust_0082 |
| GO:0008152 | metabolism | 0.815578 | 1 | Thaps_hclust_0082 |
| GO:0030261 | chromosome condensation | 0.0105437 | 1 | Thaps_hclust_0082 |
| GO:0006118 | electron transport | 0.944985 | 1 | Thaps_hclust_0082 |
| GO:0019307 | mannose biosynthesis | 0.0105437 | 1 | Thaps_hclust_0082 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.965341 | 1 | Thaps_hclust_0082 |
| GO:0000278 | mitotic cell cycle | 0.0105437 | 1 | Thaps_hclust_0082 |
|
370 : (RIR1_1) PLN02437 |
6699 : hypothetical protein |
14707 : Motor_domain superfamily |
33979 : S_TKc |
|
1111 : GINS_A_psf3 |
7091 : hypothetical protein |
15138 : MFS |
34210 : PLN00158 |
|
1224 : MFS transporter |
7092 : DOMON_DOH |
15226 : PLN02407 |
35310 : KISc_C_terminal |
|
1393 : (CAP1) Smc |
7100 : Condensin2nSMC |
19793 : PLN00157 |
35387 : (cdc2) PKc_like superfamily |
|
1422 : Cnd3 |
7206 : hypothetical protein |
20673 : PTZ00121 |
35968 : RecA-like_NTPases superfamily |
|
1622 : AE_Prim_S |
7586 : MAP65_ASE1 |
20798 : Cnd1 |
36441 : CYCLIN |
|
2267 : hypothetical protein |
8031 : DUF1028 superfamily |
20903 : CAF1A |
36753 : Motor_domain superfamily |
|
2376 : PTZ00298 |
8522 : RNRR2 |
21094 : hypothetical protein |
36788 : PLN02438 |
|
3124 : Cnd2 superfamily |
8536 : CDC6 |
22301 : (PMM1) PLN02423 |
37398 : Motor_domain superfamily |
|
4041 : HEC1 |
8829 : hypothetical protein |
23012 : hypothetical protein |
37810 : GMPK |
|
4170 : hypothetical protein |
9101 : hypothetical protein |
23028 : KISc_KIF2_like |
42365 : Smc |
|
4214 : hypothetical protein |
9684 : TIMELESS superfamily |
23064 : hypothetical protein |
260808 : |
|
4217 : hypothetical protein |
9991 : hypothetical protein |
23522 : DUF1032 superfamily |
261327 : NADB_Rossmann superfamily |
|
4607 : hypothetical protein |
9992 : H15 |
24220 : hypothetical protein |
261826 : SEC14 |
|
4688 : hypothetical protein |
9993 : H15 |
24304 : hypothetical protein |
262006 : alpha_CA_prokaryotic_like |
|
4729 : Glycos_transf_1 superfamily |
10002 : Solute_trans_a superfamily |
25898 : hypothetical protein |
262009 : alpha_CA superfamily |
|
4808 : hypothetical protein |
10156 : Tcp10_C superfamily |
25910 : hypothetical protein |
262102 : KISc |
|
5009 : hypothetical protein |
10434 : hypothetical protein |
25917 : hypothetical protein |
263366 : Pkinase |
|
5263 : hypothetical protein |
10832 : hypothetical protein |
31569 : (TUB3) PLN00220 |
263798 : incenp-like protein |
|
5431 : hypothetical protein |
11186 : hypothetical protein |
32555 : RNRR2 |
268117 : H2A |
|
5662 : hypothetical protein |
11559 : hypothetical protein |
32661 : ABC_ATPase superfamily |
268221 : (Tp_Myb1R10) MYB DNA binding protein/ transcription factor-like protein |
|
5867 : CDC6 |
11643 : (Tp_AP2-EREBP3) regulator [Rayko] |
33513 : CYCLIN |
268247 : SLBP_RNA_bind |
|
6356 : hypothetical protein |
11918 : hypothetical protein |
33794 : ERG4_ERG24 superfamily |
269422 : wd-40 repeat-containing protein |
|
6586 : hypothetical protein |
11958 : Nuf2 superfamily |
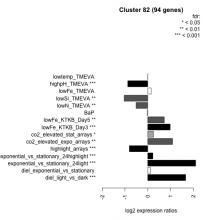
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| diel_light_vs_dark | diel_light_vs_dark | 1.670 | 0.000485 |
| lowFe_KTKB_Day3 | lowFe_KTKB_Day3 | 0.987 | 0.000862 |
| lowFe_KTKB_Day5 | lowFe_KTKB_Day5 | 0.728 | 0.00132 |
| BaP | BaP | -0.027 | 0.824 |
| exponential_vs_stationary_24highlight | exponential_vs_stationary_24highlight | 0.226 | 0.000526 |
| co2_elevated_stat_arrays | co2_elevated_stat_arrays | 0.253 | 0.0468 |
| lowtemp_TMEVA | lowtemp_TMEVA | -0.001 | 1 |
| highpH_TMEVA | highpH_TMEVA | -0.879 | 0.000725 |
| co2_elevated_expo_arrays | co2_elevated_expo_arrays | 1.090 | 0.00139 |
| lowFe_TMEVA | lowFe_TMEVA | 0.152 | 0.452 |
| exponential_vs_stationary_24light | exponential_vs_stationary_24light | 2.110 | 0.000581 |
| lowN_TMEVA | lowN_TMEVA | -0.512 | 0.00119 |
| diel_exponential_vs_stationary | diel_exponential_vs_stationary | 0.116 | 0.17 |
| lowSi_TMEVA | lowSi_TMEVA | -1.030 | 0.00135 |
| highlight_arrays | highlight_arrays | -0.807 | 0.000442 |