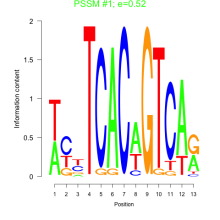
Phatr_bicluster_0044 Residual: 0.37
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0044 | 0.37 | Phaeodactylum tricornutum |
Displaying 1 - 36 of 36
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_1103 calcium-dependent protein kinase (S_TKc)
PHATRDRAFT_13320 (COG0412)
PHATRDRAFT_17720 long chain acyl-coa synthetase (PLN02736)
PHATRDRAFT_19333 fas2 (fasciata 2) nucleotide binding (WD40 superfamily)
PHATRDRAFT_31980 trna-specific adenosine deaminase (A_deamin superfamily)
PHATRDRAFT_32772 f119b_bovin protein fam119b (Methyltransf_16 superfamily)
PHATRDRAFT_33705 basic-leucine zippertranscription factor
PHATRDRAFT_34923 glutamineclass i (GATase1_1)
PHATRDRAFT_43746 (ATP-grasp_4 superfamily)
PHATRDRAFT_45017 molybdenum cofactor biosynthesis protein (MoeA)
PHATRDRAFT_45353 isocitrate dehydrogenase nadp-monomeric type (IDH superfamily)
PHATRDRAFT_45623 hypothetical conserved protein (AIM24)
PHATRDRAFT_46131 (AIM24)
PHATRDRAFT_46709 vacuolar sorting receptor-like protein (PA)
PHATRDRAFT_47531 protein serine threonine (S_TKc)
PHATRDRAFT_48675 (CAF1A)
PHATRDRAFT_50241 cip1 (cop1-interactive protein 1)
PHATRDRAFT_54863 (alpha_CA superfamily)
PHATRDRAFT_9413 epsilon frustilin
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0008152 | metabolism | 0.453158 | 1 | Phatr_bicluster_0044 |
| GO:0006777 | Mo-molybdopterin cofactor biosynthesis | 0.0118036 | 1 | Phatr_bicluster_0044 |
| GO:0006468 | protein amino acid phosphorylation | 0.00360249 | 1 | Phatr_bicluster_0044 |
| GO:0016567 | protein ubiquitination | 0.13382 | 1 | Phatr_bicluster_0044 |
| GO:0006396 | RNA processing | 0.105521 | 1 | Phatr_bicluster_0044 |
| GO:0006099 | tricarboxylic acid cycle | 0.015711 | 1 | Phatr_bicluster_0044 |


Comments