Phatr_bicluster_0071 Residual: 0.39
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0071 | 0.39 | Phaeodactylum tricornutum |
Displaying 1 - 22 of 22
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12808 copper amine oxidase (Cu_amine_oxid)
PHATRDRAFT_14294 (PITH)
PHATRDRAFT_25168 light harvesting protein (PLN00120)
PHATRDRAFT_27877 ammonium transporter (Ammonium_transp superfamily)
PHATRDRAFT_28245 chloride channel 7 (Voltage_gated_ClC superfamily)
PHATRDRAFT_35050 (Tmemb_14)
PHATRDRAFT_36006 type i secretion target repeat
PHATRDRAFT_39534 conserved hypothetical membrane protein (Bestrophin superfamily)
PHATRDRAFT_40280 carbohydrate-binding protein
PHATRDRAFT_42902 (COG0523)
PHATRDRAFT_43479 (PDZ_signaling)
PHATRDRAFT_43708 mgc82689 protein (SET)
PHATRDRAFT_43723 novel protein (TPD)
PHATRDRAFT_44673 (DUF4336 superfamily)
PHATRDRAFT_47436 5-nucleotidase 2-cyclic phosphodiesterase and related esterase (MPP_superfamily superfamily)
PHATRDRAFT_49244 cax family of cationmembrane protein (PLN03151)
PHATRDRAFT_5419 (HpcH_HpaI superfamily)
PHATRDRAFT_9280 calcineurin-like phosphoesterase family protein (MPP_superfamily superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0001682 | tRNA 5'-leader removal | 0.00123487 | 1 | Phatr_bicluster_0071 |
| GO:0006821 | chloride transport | 0.00861852 | 1 | Phatr_bicluster_0071 |
| GO:0009765 | photosynthesis light harvesting | 0.0472641 | 1 | Phatr_bicluster_0071 |
| GO:0005975 | carbohydrate metabolism | 0.0754495 | 1 | Phatr_bicluster_0071 |
| GO:0006810 | transport | 0.21473 | 1 | Phatr_bicluster_0071 |

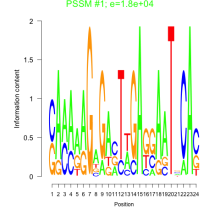
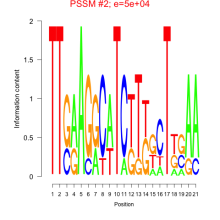
Comments