Phatr_bicluster_0084 Residual: 0.29
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0084 | 0.29 | Phaeodactylum tricornutum |
Displaying 1 - 29 of 29
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_15980 tetrapyrrole-binding protein (GUN4 superfamily)
PHATRDRAFT_16140 uroporphyrinogen decarboxylase (PLN02433)
PHATRDRAFT_16322 fucoxanthin chlorophyll a c binding protein (Chloroa_b-bind superfamily)
PHATRDRAFT_20757 uroporphyrinogen decarboxylase (URO-D)
PHATRDRAFT_22819 peroxisomal membrane (Mpv17_PMP22)
PHATRDRAFT_24119 chloroplast light harvesting protein isoform 5 (Chloroa_b-bind)
PHATRDRAFT_30690 isoflavone reductase (PLN02657)
PHATRDRAFT_31109 plastid protoporphyrinogen oxidase (PLN02576)
PHATRDRAFT_33017 mg chelatase subunit (Cob-chelat-sub)
PHATRDRAFT_36103 (TLD superfamily)
PHATRDRAFT_36362 peptidylprolyl isomerase (FKBP_C)
PHATRDRAFT_40199 (DUF1995 superfamily)
PHATRDRAFT_41515 enolase 2 (PLN00191)
PHATRDRAFT_44056 precursor cytochrome c6 (Cytochrom_C superfamily)
PHATRDRAFT_45339 (SNc)
PHATRDRAFT_48328 carbohydratefamily 1 (Esterase)
PHATRDRAFT_48875 la_aedalla protein homolog (la ribonucleoprotein) (la autoantigen homolog) (LAM)
PHATRDRAFT_49639 taurine catabolism dioxygenase (TauD)
PHATRDRAFT_50705 light harvesting protein (PLN00120)
PHATRDRAFT_55046 ests gb
PHATRDRAFT_7626 fkbp-type peptidyl-prolyl cis-trans isomerase (FKBP_C)
PHATRDRAFT_8324 cog1359: uncharacterized conserved protein (ABM superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006779 | porphyrin biosynthesis | 0.00000389 | 0.00902469 | Phatr_bicluster_0084 |
| GO:0009765 | photosynthesis light harvesting | 0.000279566 | 0.640765 | Phatr_bicluster_0084 |
| GO:0015995 | chlorophyll biosynthesis | 0.0103397 | 1 | Phatr_bicluster_0084 |
| GO:0015979 | photosynthesis | 0.0137641 | 1 | Phatr_bicluster_0084 |
| GO:0017000 | antibiotic biosynthesis | 0.0171775 | 1 | Phatr_bicluster_0084 |
| GO:0006457 | protein folding | 0.0600726 | 1 | Phatr_bicluster_0084 |
| GO:0006096 | glycolysis | 0.114621 | 1 | Phatr_bicluster_0084 |
| GO:0006118 | electron transport | 0.185759 | 1 | Phatr_bicluster_0084 |

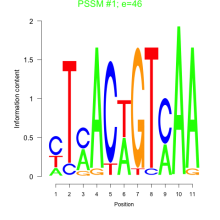
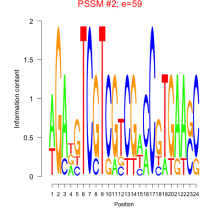
Comments