Phatr_bicluster_0186 Residual: 0.24
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0186 | 0.24 | Phaeodactylum tricornutum |
Displaying 1 - 24 of 24
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12121 atath13 (abc2 homolog 13) (AarF)
PHATRDRAFT_12663 lactoylglutathione lyase (Glyoxalase_I)
PHATRDRAFT_14125 monogalactosyldiacylglycerol synthase (PLN02605)
PHATRDRAFT_14307 possible ferredoxin (2fe-2s)
PHATRDRAFT_26077 d-isomer specific 2-hydroxyacidnad-binding (2-Hacid_dh_1)
PHATRDRAFT_31525 aspartyl protease family protein (pepsin_retropepsin_like superfamily)
PHATRDRAFT_32733 peptidase u34 dipeptidase (Peptidase_C69 superfamily)
PHATRDRAFT_32841 sel1 domain protein repeat-containing protein (COG0790)
PHATRDRAFT_34060 pentatricopeptiderepeat-containing protein
PHATRDRAFT_36844 tpr repeat
PHATRDRAFT_38508 (DUF1995 superfamily)
PHATRDRAFT_43140 arylsulfatase b (AslA)
PHATRDRAFT_43177 voltage-gated shaker-like k+subunit beta kcnab (Aldo_ket_red)
PHATRDRAFT_44306 set and mynd domain containing protein
PHATRDRAFT_44775 protein phosphatase 2c (PP2Cc)
PHATRDRAFT_45431 loc100036796 protein (G_glu_transpept superfamily)
PHATRDRAFT_45830 histone deacetylase superfamily (Arginase_HDAC superfamily)
PHATRDRAFT_46084 (NADB_Rossmann superfamily)
PHATRDRAFT_48791 arystal structuremidohydrolase bh0493 from bacillus halodurans c-125 (UxaC superfamily)
PHATRDRAFT_54536 thiol-disulphide oxidoreductase dcc (DUF393)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006564 | L-serine biosynthesis | 0.0234852 | 1 | Phatr_bicluster_0186 |
| GO:0015992 | proton transport | 0.0407661 | 1 | Phatr_bicluster_0186 |
| GO:0051341 | regulation of oxidoreductase activity | 0.0464638 | 1 | Phatr_bicluster_0186 |
| GO:0008152 | metabolism | 0.051036 | 1 | Phatr_bicluster_0186 |
| GO:0009058 | biosynthesis | 0.125669 | 1 | Phatr_bicluster_0186 |
| GO:0006508 | proteolysis and peptidolysis | 0.13969 | 1 | Phatr_bicluster_0186 |
| GO:0005975 | carbohydrate metabolism | 0.171749 | 1 | Phatr_bicluster_0186 |
| GO:0006468 | protein amino acid phosphorylation | 0.406444 | 1 | Phatr_bicluster_0186 |
| GO:0006118 | electron transport | 0.503223 | 1 | Phatr_bicluster_0186 |

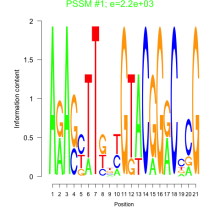

Comments