Phatr_bicluster_0217 Residual: 0.19
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0217 | 0.19 | Phaeodactylum tricornutum |
Displaying 1 - 24 of 24
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_11006 light-harvesting protein (Chloroa_b-bind)
PHATRDRAFT_14442 light-harvesting protein (Chloroa_b-bind superfamily)
PHATRDRAFT_17326 low molecular mass early light-inducible protein hv60
PHATRDRAFT_17531 fucoxanthin chlorophyll a c binding protein (Chloroa_b-bind)
PHATRDRAFT_17766 chloroplast light harvesting protein isoform 12 (Chloroa_b-bind superfamily)
PHATRDRAFT_18049 chlorophyll protein 1 (PLN00120)
PHATRDRAFT_20331 oxygen-evolving enhancer protein 1 precursor (MSP superfamily)
PHATRDRAFT_22006 fucoxanthin chlorophyll protein 3 (PLN00120)
PHATRDRAFT_22680 chloroplast light harvesting protein isoform 15 (Chloroa_b-bind superfamily)
PHATRDRAFT_22956 light-harvesting protein (Chloroa_b-bind)
PHATRDRAFT_23257 chloroplast light harvesting protein isoform 14 (Chloroa_b-bind superfamily)
PHATRDRAFT_25172 fcpb_phatrfucoxanthin-chlorophyll a-c binding proteinchloroplast precursor (PLN00120)
PHATRDRAFT_25893 light-harvesting chlorophyll a-c binding protein (Chloroa_b-bind superfamily)
PHATRDRAFT_26293 chloroplast photosystem ii 12 kda extrinsic protein (PsbU)
PHATRDRAFT_29266 fcpb_phatrfucoxanthin-chlorophyll a-c binding proteinchloroplast precursor (PLN00120)
PHATRDRAFT_30031 light harvesting protein (PLN00120)
PHATRDRAFT_30643 fcpb_phatrfucoxanthin-chlorophyll a-c binding proteinchloroplast precursor (PLN00120)
PHATRDRAFT_30648 fucoxanthin chlorophyll protein 3 (PLN00120)
PHATRDRAFT_51230 fcpf_phatrfucoxanthin-chlorophyll a-c binding proteinchloroplast precursor (PLN00120)
PHATRDRAFT_54027 light-harvesting protein (Chloroa_b-bind superfamily)
PHATRDRAFT_6062 light-harvesting protein (Chloroa_b-bind)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0009765 | photosynthesis light harvesting | 5.21e-33 | 1.21e-29 | Phatr_bicluster_0217 |
| GO:0042549 | photosystem II stabilization | 0.0000256 | 0.0592446 | Phatr_bicluster_0217 |
| GO:0015979 | photosynthesis | 0.0205926 | 1 | Phatr_bicluster_0217 |
| GO:0007186 | G-protein coupled receptor protein signaling pathway | 0.0654593 | 1 | Phatr_bicluster_0217 |

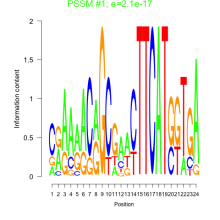
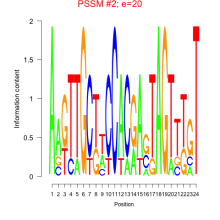
Comments