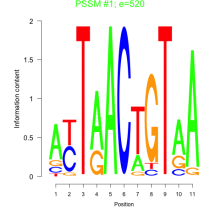
Phatr_bicluster_0221 Residual: 0.32
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0221 | 0.32 | Phaeodactylum tricornutum |
Displaying 1 - 29 of 29
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_17265 proton phosphate symporter (MFS)
PHATRDRAFT_31257 dihydrokaempferol 4-reductase (NADB_Rossmann superfamily)
PHATRDRAFT_36256 acetate kinase (Acetate_kinase superfamily)
PHATRDRAFT_42561 histidine acid phosphatase family protein (HP_HAP_like)
PHATRDRAFT_43118 hormone sensitive (Abhydrolase_3)
PHATRDRAFT_44525 (SET)
PHATRDRAFT_44809 chromosome 3 open reading frame 37 (DUF159)
PHATRDRAFT_45141 3-hydroxyisobutyrate dehydrogenase (HIBADH)
PHATRDRAFT_45728 at3g52570 f22o6_50 (MhpC)
PHATRDRAFT_45755 taurine catabolism dioxygenasefamily (CAS_like superfamily)
PHATRDRAFT_45982 cg13101-isoform a
PHATRDRAFT_45986 flagellar calcium-binding (EFh)
PHATRDRAFT_47077 family protein (NADB_Rossmann superfamily)
PHATRDRAFT_47141 zinc-binding oxidoreductase (MDR_like_2)
PHATRDRAFT_49064 (NhaC)
PHATRDRAFT_49251 cral trio domain containing protein (SEC14)
PHATRDRAFT_50281 fumarylacetoacetase (PLN02856)
PHATRDRAFT_51609 tyrosine aminotransferase (tyr_nico_aTase)
PHATRDRAFT_54560 nitrate transporter (2A0108)
PHATRDRAFT_6848 maleylacetoacetate isomerase (Thioredoxin_like superfamily)
PHATRDRAFT_843 methylcrotonoyl-carboxylase alphamitochondrialexpressed (PRK08591)
PHATRDRAFT_9476 2-oxoisovalerate dehydrogenase alphamitochondrialexpressed (TPP_E1_PDC_ADC_BCADC)
PHATRDRAFT_9728 carbon-sulfur lyase (GFA superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0008152 | metabolism | 0.00154435 | 1 | Phatr_bicluster_0221 |
| GO:0006082 | organic acid metabolism | 0.00321067 | 1 | Phatr_bicluster_0221 |
| GO:0009072 | aromatic amino acid family metabolism | 0.00960348 | 1 | Phatr_bicluster_0221 |
| GO:0006573 | valine metabolism | 0.00960348 | 1 | Phatr_bicluster_0221 |
| GO:0006098 | pentose-phosphate shunt | 0.0191218 | 1 | Phatr_bicluster_0221 |
| GO:0006885 | regulation of pH | 0.0254203 | 1 | Phatr_bicluster_0221 |
| GO:0006814 | sodium ion transport | 0.0347981 | 1 | Phatr_bicluster_0221 |
| GO:0016310 | phosphorylation | 0.0440926 | 1 | Phatr_bicluster_0221 |
| GO:0006810 | transport | 0.466944 | 1 | Phatr_bicluster_0221 |


Comments