Phatr_bicluster_0248 Residual: 0.29
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0248 | 0.29 | Phaeodactylum tricornutum |
Displaying 1 - 31 of 31
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12292 ribosomal protein l33 (rpmG)
PHATRDRAFT_12378 subfamilymember 9 (DnaJ)
PHATRDRAFT_12921 alm beta-like (S1_like superfamily)
PHATRDRAFT_13160 farnesyl diphosphate farnesyl transferase 1 (Trans_IPPS_HH)
PHATRDRAFT_13167 solute carrier family 35 (UAA)
PHATRDRAFT_18036 dnaj domain (DnaJ)
PHATRDRAFT_20120 solute carrier family 35 (udp-n-acetylglucosamine (udp-c) transporter)memberisoform cra_b (Nuc_sug_transp)
PHATRDRAFT_20508 long chain polyunsaturated fatty acid elongation (ELO)
PHATRDRAFT_21275 (Abhydrolase_5)
PHATRDRAFT_27735 gmp synthase (PLN02347)
PHATRDRAFT_28551 dna-dependent atpase snf2h (PLN03142)
PHATRDRAFT_31662 mono- or diacylglycerol acyltransferase (LPLAT superfamily)
PHATRDRAFT_31927 peroxisomal protein pex19 family protein (Pex19 superfamily)
PHATRDRAFT_41875 3-bisphosphate nucleotidase (FIG superfamily)
PHATRDRAFT_42673 sec23_ustma protein transport protein sec23 (PLN00162)
PHATRDRAFT_42721 conserved protein (Gly_transf_sug superfamily)
PHATRDRAFT_43774 formin 2 (Nup54 superfamily)
PHATRDRAFT_44312 (SMC_prok_B)
PHATRDRAFT_44505 aspartyl asparaginyl beta- (Asp_Arg_Hydrox superfamily)
PHATRDRAFT_44547 ankyrin repeat (Pho88 superfamily)
PHATRDRAFT_44858 spc25_schpo kinetochore protein spc25 (Spindle_Spc25)
PHATRDRAFT_45621 hmp_bacsu flavohemoprotein (hemoglobin-like protein)(nitric oxide dioxygenase) (no oxygenase) (PRK13289)
PHATRDRAFT_45650 nuclear pore protein (RanBD_family)
PHATRDRAFT_46021 serine threonine rich antigen
PHATRDRAFT_46074 aspartyl asparaginyl beta-hydroxylase (Asp_Arg_Hydrox superfamily)
PHATRDRAFT_47054 taf12 rna polymerasetata box binding protein-associated factor (TAF12)
PHATRDRAFT_48517 sam-dependent methyltransferases (Cfa)
PHATRDRAFT_9123 gcn4-complementing protein homolog (ArfGap superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0008610 | lipid biosynthesis | 0.0000693 | 0.159914 | Phatr_bicluster_0248 |
| GO:0018193 | peptidyl-amino acid modification | 0.000229788 | 0.527133 | Phatr_bicluster_0248 |
| GO:0006177 | GMP biosynthesis | 0.0098558 | 1 | Phatr_bicluster_0248 |
| GO:0006164 | purine nucleotide biosynthesis | 0.0196193 | 1 | Phatr_bicluster_0248 |
| GO:0015780 | nucleotide-sugar transport | 0.0196193 | 1 | Phatr_bicluster_0248 |
| GO:0009058 | biosynthesis | 0.0202116 | 1 | Phatr_bicluster_0248 |
| GO:0006888 | ER to Golgi transport | 0.0388725 | 1 | Phatr_bicluster_0248 |
| GO:0043087 | regulation of GTPase activity | 0.0530761 | 1 | Phatr_bicluster_0248 |
| GO:0006352 | transcription initiation | 0.0577662 | 1 | Phatr_bicluster_0248 |
| GO:0006633 | fatty acid biosynthesis | 0.0624342 | 1 | Phatr_bicluster_0248 |

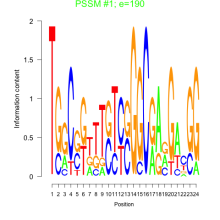
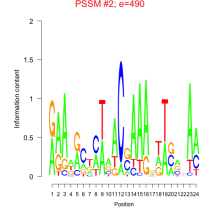
Comments