261967 (Tp_Hox7) regulator [Rayko]Thalassiosira pseudonana
| Chromosome | Product | Transcript Start | End | Strand | Short Name | |
|---|---|---|---|---|---|---|
| 261967 | chr_3 | (Tp_Hox7) regulator [Rayko] | 2282287 | 2282661 | + | regulator [Rayko] |
| NCBI ID | Ensembl Genomes exon ID |
|---|---|
| 7443821 | Thaps261967.1 |
| Expression Profile | Conditional Changes | Cluster Dendrogram | Discovered Potential cis-Regulatory Motifs |
|---|---|---|---|
Thaps_hclust_0353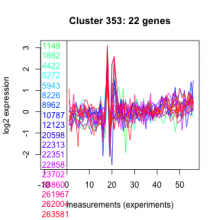 |
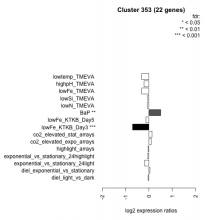 |
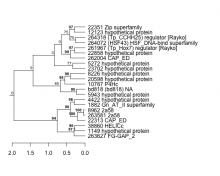 |
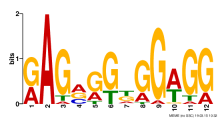 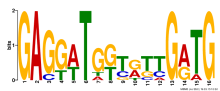  |
| Normalized Mean Residue | Discovered Potential cis-Regulatory Motifs | |
|---|---|---|
|
Thaps_bicluster_0026 |
0.39 |
Not available
| KEGG description | KEGG Pathway |
|---|---|
| Not available | Not available |

Add comment