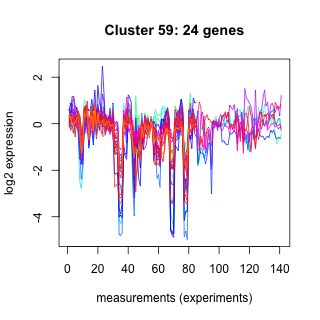
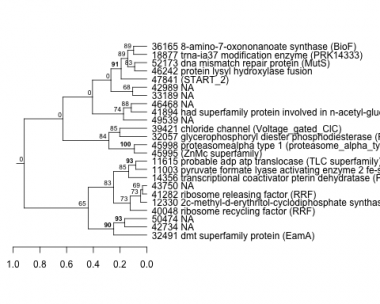
Phatr_hclust_0059 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0016114 | terpenoid biosynthesis | 0.004934 | 1 | Phatr_hclust_0059 |
| GO:0006071 | glycerol metabolism | 0.00984608 | 1 | Phatr_hclust_0059 |
| GO:0006821 | chloride transport | 0.0171733 | 1 | Phatr_hclust_0059 |
| GO:0006412 | protein biosynthesis | 0.0573335 | 1 | Phatr_hclust_0059 |
| GO:0006511 | ubiquitin-dependent protein catabolism | 0.105854 | 1 | Phatr_hclust_0059 |
| GO:0006810 | transport | 0.38354 | 1 | Phatr_hclust_0059 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.426896 | 1 | Phatr_hclust_0059 |
| GO:0006508 | proteolysis and peptidolysis | 0.434343 | 1 | Phatr_hclust_0059 |
| GO:0008152 | metabolism | 0.529843 | 1 | Phatr_hclust_0059 |
|
PHATRDRAFT_32491 : dmt superfamily protein (EamA) |
PHATRDRAFT_32057 : glycerophosphoryl diester phosphodiesterase (PI-PLCc_GDPD_SF superfamily) |
||
|
PHATRDRAFT_14356 : transcriptional coactivator pterin dehydratase (PCD_DCoH superfamily) |
PHATRDRAFT_39421 : chloride channel (Voltage_gated_ClC) |
PHATRDRAFT_47841 : (START_2) |
|
|
PHATRDRAFT_11003 : pyruvate formate lyase activating enzyme 2 fe-s cluster domain (Radical_SAM) |
PHATRDRAFT_46242 : protein lysyl hydroxylase fusion |
||
|
PHATRDRAFT_40048 : ribosome recycling factor (RRF) |
PHATRDRAFT_11615 : probable adp atp translocase (TLC superfamily) |
PHATRDRAFT_41894 : had superfamily protein involved in n-acetyl-glucosamine catabolism-like (HAD-SF-IIA-hyp4) |
PHATRDRAFT_52173 : dna mismatch repair protein (MutS) |
|
PHATRDRAFT_12330 : 2c-methyl-d-erythritol-cyclodiphosphate synthase (MECDP_synthase) |
PHATRDRAFT_45995 : (ZnMc superfamily) |
PHATRDRAFT_18877 : trna-ia37 modification enzyme (PRK14333) |
|
|
PHATRDRAFT_41282 : ribosome releasing factor (RRF) |
PHATRDRAFT_45998 : proteasomealpha type 1 (proteasome_alpha_type_1) |
PHATRDRAFT_36165 : 8-amino-7-oxononanoate synthase (BioF) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_6h | Re-illuminated_6h | 0.023 | 0.934277 |
| Copper_SH | Copper_SH | 0.015 | 0.8842 |
| Green_vs_Red_24h | Green_vs_Red_24h | 0.070 | 0.557285 |
| Silver_SH | Silver_SH | 0.203 | 0.0301932 |
| Cadmium_1.2mg | Cadmium_1.2mg | 0.014 | 0.908681 |
| BlueLight_24h | BlueLight_24h | 0.094 | 0.701384 |
| GreenLight_0.5h | GreenLight_0.5h | -2.136 | 0.000458716 |
| RedLight_6h | RedLight_6h | -0.218 | 0.306522 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 1 |
| GreenLight_24h | GreenLight_24h | 0.045 | 0.850369 |
| light_6hr | light_6hr | -0.692 | 0.00229358 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | -0.454 | 0.0194245 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.037 | 0.776956 |
| Ammonia_SH | Ammonia_SH | -0.710 | 0.00820513 |
| Simazine_SH | Simazine_SH | 0.011 | 0.921677 |
| Oil_SH | Oil_SH | -0.113 | 0.25307 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | 0.191 | 0.0936567 |
| Cadmium_SH | Cadmium_SH | -0.735 | 0.000666667 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | 0.119 | 0.399436 |
| highlight_0to6h | highlight_0to6h | -0.166 | 0.223515 |
| BlueLight_0.5h | BlueLight_0.5h | -2.213 | 0.000462963 |
| Dispersant_SH | Dispersant_SH | 0.012 | 0.92463 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.148 | 0.204887 |
| light_15.5hr | light_15.5hr | -0.387 | 0.0960938 |
| light_10.5hr | light_10.5hr | -0.877 | 0.000561798 |
| Dispersed_oil_SH | Dispersed_oil_SH | -0.056 | 0.589186 |
| Si_free | Si_free | 0.241 | 0.15875 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | -2.366 | 0.000396825 |
| Mixture_SH | Mixture_SH | -0.151 | 0.611833 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | 0.229 | 0.240741 |
| light_16hr_dark_30min | light_16hr_dark_30min | -0.133 | 0.615173 |
| BlueLight_6h | BlueLight_6h | 0.012 | 0.965409 |
| highlight_12to48h | highlight_12to48h | 0.218 | 0.254412 |
| RedLight_24h | RedLight_24h | -0.026 | 0.886403 |
| Dark_treated | Dark_treated | -0.753 | 0.186275 |
| GreenLight_6h | GreenLight_6h | -0.025 | 0.898113 |
| Cadmium_0.12mg | Cadmium_0.12mg | 0.049 | 0.22226 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | 0.049 | 0.55057 |
| RedLight_0.5h | RedLight_0.5h | -0.924 | 0.0104478 |
| Green_vs_Red_6h | Green_vs_Red_6h | 0.193 | 0.300287 |
| Re-illuminated_24h | Re-illuminated_24h | 0.170 | 0.205725 |