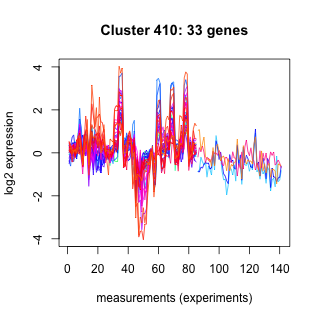
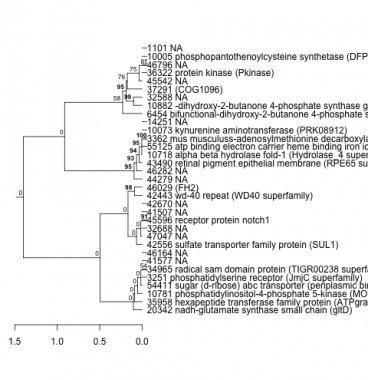
Phatr_hclust_0410 Hierarchical Clustering
Phaeodactylum tricornutum
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0009231 | riboflavin biosynthesis | 0.000141735 | 0.445331 | Phatr_hclust_0410 |
| GO:0008295 | spermidine biosynthesis | 0.00321067 | 1 | Phatr_hclust_0410 |
| GO:0006597 | spermine biosynthesis | 0.00321067 | 1 | Phatr_hclust_0410 |
| GO:0016043 | cell organization and biogenesis | 0.00641182 | 1 | Phatr_hclust_0410 |
| GO:0006537 | glutamate biosynthesis | 0.0159584 | 1 | Phatr_hclust_0410 |
| GO:0009058 | biosynthesis | 0.135415 | 1 | Phatr_hclust_0410 |
| GO:0007165 | signal transduction | 0.141021 | 1 | Phatr_hclust_0410 |
| GO:0005975 | carbohydrate metabolism | 0.184674 | 1 | Phatr_hclust_0410 |
| GO:0006468 | protein amino acid phosphorylation | 0.431733 | 1 | Phatr_hclust_0410 |
| GO:0006508 | proteolysis and peptidolysis | 0.523353 | 1 | Phatr_hclust_0410 |
|
PHATRDRAFT_20342 : nadh-glutamate synthase small chain (gltD) |
PHATRDRAFT_42556 : sulfate transporter family protein (SUL1) |
PHATRDRAFT_10882 : -dihydroxy-2-butanone 4-phosphate synthase gtp cyclohydrolase ii (GTP_cyclohydro2) |
|
|
PHATRDRAFT_35958 : hexapeptide transferase family protein (ATPgrasp_ST superfamily) |
|||
|
PHATRDRAFT_10781 : phosphatidylinositol-4-phosphate 5-kinase (MORN superfamily) |
PHATRDRAFT_43490 : retinal pigment epithelial membrane (RPE65 superfamily) |
PHATRDRAFT_37291 : (COG1096) |
|
|
PHATRDRAFT_54411 : sugar (d-ribose) abc transporter (periplasmic binding protein) (Peripla_BP_4) |
PHATRDRAFT_45596 : receptor protein notch1 |
PHATRDRAFT_10718 : alpha beta hydrolase fold-1 (Hydrolase_4 superfamily) |
|
|
PHATRDRAFT_3251 : phosphatidylserine receptor (JmjC superfamily) |
PHATRDRAFT_3362 : mus musculuss-adenosylmethionine decarboxylase 1 (PLN02524) |
PHATRDRAFT_36322 : protein kinase (Pkinase) |
|
|
PHATRDRAFT_34965 : radical sam domain protein (TIGR00238 superfamily) |
PHATRDRAFT_10073 : kynurenine aminotransferase (PRK08912) |
||
|
PHATRDRAFT_42443 : wd-40 repeat (WD40 superfamily) |
PHATRDRAFT_10005 : phosphopantothenoylcysteine synthetase (DFP superfamily) |
||
|
PHATRDRAFT_46029 : (FH2) |
PHATRDRAFT_6454 : bifunctional-dihydroxy-2-butanone 4-phosphate synthase gtp cyclohydrolase ii protein (DHBP_synthase) |
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| Re-illuminated_6h | Re-illuminated_6h | 0.096 | 0.647407 |
| Copper_SH | Copper_SH | -0.797 | 0.000515464 |
| Green_vs_Red_24h | Green_vs_Red_24h | -0.093 | 0.359053 |
| Silver_SH | Silver_SH | -0.728 | 0.000555556 |
| Cadmium_1.2mg | Cadmium_1.2mg | -0.042 | 0.558893 |
| BlueLight_24h | BlueLight_24h | 0.229 | 0.0489474 |
| GreenLight_0.5h | GreenLight_0.5h | 1.494 | 0.000458716 |
| RedLight_6h | RedLight_6h | 0.158 | 0.410213 |
| light_16hr_dark_7.5hr | light_16hr_dark_7.5hr | 0.000 | 0.980594 |
| GreenLight_24h | GreenLight_24h | 0.321 | 0.000961538 |
| light_6hr | light_6hr | -0.964 | 0.000561798 |
| dark_8hr_light_3hr | dark_8hr_light_3hr | -0.773 | 0.00060241 |
| Blue_vs_Green_6h | Blue_vs_Green_6h | 0.089 | 0.28526 |
| Ammonia_SH | Ammonia_SH | -0.135 | 0.55615 |
| Simazine_SH | Simazine_SH | -0.851 | 0.000423729 |
| Oil_SH | Oil_SH | -0.257 | 0.00905512 |
| Salinity_15ppt_SH | Salinity_15ppt_SH | -0.647 | 0.000609756 |
| Cadmium_SH | Cadmium_SH | -0.509 | 0.000666667 |
| Blue_vs_Red_24h | Blue_vs_Red_24h | -0.185 | 0.153378 |
| highlight_0to6h | highlight_0to6h | 0.263 | 0.00523256 |
| BlueLight_0.5h | BlueLight_0.5h | 1.764 | 0.000462963 |
| Dispersant_SH | Dispersant_SH | -0.540 | 0.000909091 |
| light_16hr_dark_4hr | light_16hr_dark_4hr | -0.451 | 0.000943396 |
| light_15.5hr | light_15.5hr | -1.112 | 0.000625 |
| light_10.5hr | light_10.5hr | -1.274 | 0.000561798 |
| Dispersed_oil_SH | Dispersed_oil_SH | -0.354 | 0.000675676 |
| Si_free | Si_free | 0.096 | 0.392025 |
| Re-illuminated_0.5h | Re-illuminated_0.5h | 1.988 | 0.000396825 |
| Mixture_SH | Mixture_SH | -0.288 | 0.237042 |
| Blue_vs_Red_6h | Blue_vs_Red_6h | -0.132 | 0.471652 |
| light_16hr_dark_30min | light_16hr_dark_30min | -0.925 | 0.000735294 |
| BlueLight_6h | BlueLight_6h | 0.025 | 0.899007 |
| highlight_12to48h | highlight_12to48h | 0.328 | 0.030226 |
| RedLight_24h | RedLight_24h | 0.414 | 0.000943396 |
| Dark_treated | Dark_treated | 0.637 | 0.197701 |
| GreenLight_6h | GreenLight_6h | -0.064 | 0.70665 |
| Cadmium_0.12mg | Cadmium_0.12mg | -0.070 | 0.0292208 |
| Blue_vs_Green_24h | Blue_vs_Green_24h | -0.092 | 0.145902 |
| RedLight_0.5h | RedLight_0.5h | 1.287 | 0.000555556 |
| Green_vs_Red_6h | Green_vs_Red_6h | -0.222 | 0.154274 |
| Re-illuminated_24h | Re-illuminated_24h | 0.399 | 0.000657895 |