Phatr_bicluster_0092 Residual: 0.51
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0092 | 0.51 | Phaeodactylum tricornutum |
Displaying 1 - 55 of 55
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_22453 dass family transporter: sodium ion sulfate (ArsB_NhaD_permease superfamily)
PHATRDRAFT_33107 (Peptidase_M11 superfamily)
PHATRDRAFT_37811 (PEX11 superfamily)
PHATRDRAFT_38097 (HMGB-UBF_HMG-box)
PHATRDRAFT_38100 chondroitin sulfate proteoglycan 2
PHATRDRAFT_38549 guanylyl cyclase family member (gcy-28) (CHD)
PHATRDRAFT_39785 heat shock transcription factor 4 isoform 1 (HSF_DNA-bind)
PHATRDRAFT_39880 high mobility group protein 4 (hmg-4) (high mobility group protein 2a) (hmg-2a) (HMGB-UBF_HMG-box)
PHATRDRAFT_40473 copia protein (gag-int-pol protein)
PHATRDRAFT_40896 (ANK)
PHATRDRAFT_41130 cog0398: uncharacterized conserved protein (COG0398)
PHATRDRAFT_41508 cobalamin synthesis protein p47k family protein (COG0523)
PHATRDRAFT_47550 sarcosine oxidase (NAD_binding_8 superfamily)
PHATRDRAFT_47560 iesterase family member (pde-4)
PHATRDRAFT_47737 arginine serine-rich splicing factor (RRM_Srp1p_AtRSp31_like)
PHATRDRAFT_47923 (Exostosin)
PHATRDRAFT_47958 2fe-2s iron-sulfur cluster
PHATRDRAFT_47987 proline rich protein
PHATRDRAFT_48291 calcium-dependent protein kinase (EFh)
PHATRDRAFT_48394 heat shock protein (HSF_DNA-bind)
PHATRDRAFT_48701 bzip transcription factor family protein
PHATRDRAFT_48806 start domain-containing protein (DUF1336 superfamily)
PHATRDRAFT_48881 glucosaminyl (n-acetyl) transferasecore 2
PHATRDRAFT_49025 mgc157129 protein (STKc_MAK_like)
PHATRDRAFT_49215 (vWFA superfamily)
PHATRDRAFT_49952 cell surface flocculin
PHATRDRAFT_49988 domain containing (TraB superfamily)
PHATRDRAFT_50299 novel protease (XendoU)
PHATRDRAFT_52648 beta-ketoacyl synthase (PRK07967)
PHATRDRAFT_54954 calmodulin-domain protein kinase cdpk isoform 9 (EFh)
PHATRDRAFT_55029 carbonate dehydratase (alpha_CA)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006355 | regulation of transcription, DNA-dependent | 0.0000775 | 0.178669 | Phatr_bicluster_0092 |
| GO:0005978 | glycogen biosynthesis | 0.00739642 | 1 | Phatr_bicluster_0092 |
| GO:0009190 | cyclic nucleotide biosynthesis | 0.0612637 | 1 | Phatr_bicluster_0092 |
| GO:0006813 | potassium ion transport | 0.0717043 | 1 | Phatr_bicluster_0092 |
| GO:0006979 | response to oxidative stress | 0.0854566 | 1 | Phatr_bicluster_0092 |
| GO:0007242 | intracellular signaling cascade | 0.145022 | 1 | Phatr_bicluster_0092 |
| GO:0007165 | signal transduction | 0.160913 | 1 | Phatr_bicluster_0092 |
| GO:0006508 | proteolysis and peptidolysis | 0.199918 | 1 | Phatr_bicluster_0092 |
| GO:0006468 | protein amino acid phosphorylation | 0.479142 | 1 | Phatr_bicluster_0092 |
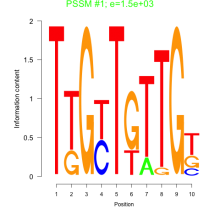


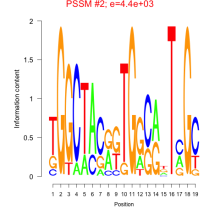
Comments