Phatr_bicluster_0119 Residual: 0.39
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0119 | 0.39 | Phaeodactylum tricornutum |
Displaying 1 - 32 of 32
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12368 myb domain-containing protein (HELICc)
PHATRDRAFT_13890 glutamyl-trnaa subunit (Amidase superfamily)
PHATRDRAFT_15323 alpha beta fold family (Abhydrolase_6)
PHATRDRAFT_21201 udp-sulfoquinovose synthase (PLN02572)
PHATRDRAFT_25183 carbamoyl-phosphate synthetaseaspartateand dihydroorotase (PLN02527)
PHATRDRAFT_28585 af193846_1branched-chain amino acid aminotransferase (PLPDE_IV superfamily)
PHATRDRAFT_30446 chaperonin containingsubunit 4 (TCP1_delta)
PHATRDRAFT_32847 glycine cleavage system h protein (GCS_H)
PHATRDRAFT_38736 (DUF1499 superfamily)
PHATRDRAFT_38794 (Glyco_hydrolase_16 superfamily)
PHATRDRAFT_39209 (SET)
PHATRDRAFT_40349 sec63 cg8583-pa (DnaJ)
PHATRDRAFT_40650 nicotinatenucleotide adenylyltransferase (NMNAT)
PHATRDRAFT_41016 aspartyl-trna synthetase (aspC)
PHATRDRAFT_42866 oxa12_schpo inner membrane protein oxa1-mitochondrial precursor (cytochrome oxidase biogenesis protein 1-2) (60KD_IMP)
PHATRDRAFT_43499 tumor suppressor candidate 4 (NPR2 superfamily)
PHATRDRAFT_44623 (YL1_C)
PHATRDRAFT_47829 rieske (2fe-2s) region (Rieske superfamily)
PHATRDRAFT_48391 jmj4_schpo jumonji domain-containing protein 4 (meiotically up-regulated gene 149 protein) (Cupin_8)
PHATRDRAFT_48637 amino acid transporter (Aa_trans superfamily)
PHATRDRAFT_49487 mgc80283 protein (FtsJ)
PHATRDRAFT_49522 cog3195: uncharacterized protein conserved in bacteria (OHCU_decarbox superfamily)
PHATRDRAFT_49610 (BglC)
PHATRDRAFT_50177 (Glyco_hydrolase_16 superfamily)
PHATRDRAFT_50496 hypothetical sam-dependent methyltransferase
PHATRDRAFT_51407 polgamma1 (polymerase gamma 1) dna binding dna-directed dna polymerase (DNA_pol_A_plastid_like)
PHATRDRAFT_7763 flag-tagged protein kinase domain ofmitogen-activated protein kinase kinase kinase (PKc_like superfamily)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006457 | protein folding | 0.410758 | 1 | Phatr_bicluster_0119 |
| GO:0006468 | protein amino acid phosphorylation | 0.542981 | 1 | Phatr_bicluster_0119 |
| GO:0006118 | electron transport | 0.65014 | 1 | Phatr_bicluster_0119 |
| GO:0006546 | glycine catabolism | 0.0132807 | 1 | Phatr_bicluster_0119 |
| GO:0051205 | protein insertion into membrane | 0.0176704 | 1 | Phatr_bicluster_0119 |
| GO:0006207 | 'de novo' pyrimidine base biosynthesis | 0.0176704 | 1 | Phatr_bicluster_0119 |
| GO:0006422 | aspartyl-tRNA aminoacylation | 0.0263947 | 1 | Phatr_bicluster_0119 |
| GO:0044267 | cellular protein metabolism | 0.0436241 | 1 | Phatr_bicluster_0119 |
| GO:0006725 | aromatic compound metabolism | 0.0854465 | 1 | Phatr_bicluster_0119 |
| GO:0006520 | amino acid metabolism | 0.101687 | 1 | Phatr_bicluster_0119 |

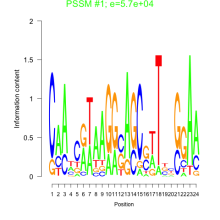
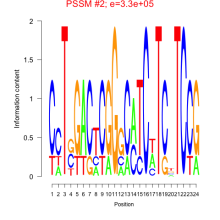
Comments