Phatr_bicluster_0165 Residual: 0.45
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0165 | 0.45 | Phaeodactylum tricornutum |
Displaying 1 - 34 of 34
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_1389 disease resistance protein
PHATRDRAFT_20948 cytochrome c isoform 1 (Cytochrom_C superfamily)
PHATRDRAFT_32294 light-harvesting protein (Chloroa_b-bind)
PHATRDRAFT_34431 (LRR_RI superfamily)
PHATRDRAFT_36711 spentranscriptional regulator
PHATRDRAFT_37567 family with sequence similaritymember a (DUF842 superfamily)
PHATRDRAFT_41570 peptidase family s49 protein (S49_Sppa_N_C)
PHATRDRAFT_43498 fatty acid desaturase family member (fat-2) (Membrane-FADS-like superfamily)
PHATRDRAFT_44232 tethering factor sec34
PHATRDRAFT_44994 atp-dependent clp protease (S14_ClpP_2)
PHATRDRAFT_45238 dnaj heat shock n-terminal domain-containing protein (DnaJ)
PHATRDRAFT_45527 thrombospondin type 3 repeat:cna b-type
PHATRDRAFT_46165 uncharacterized conserved protein (SPOUT_MTase superfamily)
PHATRDRAFT_47099 pomp cg9324-pa (UMP1 superfamily)
PHATRDRAFT_47766 conserved protein having 3 transmembrane domains and possible er retention motif possible er protein (Sec62 superfamily)
PHATRDRAFT_48128 chromosome segregation atpase
PHATRDRAFT_48867 tpr domain containing protein (TPR)
PHATRDRAFT_50191 serine arginine repetitive matrix 1
PHATRDRAFT_54019 nucleosomal binding protein
PHATRDRAFT_54217 heat shock protein 70 (PTZ00009)
PHATRDRAFT_54465 fucoxanthin chlorophyll a c binding protein (Chloroa_b-bind)
PHATRDRAFT_9547 nucleosome binding protein
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006636 | fatty acid desaturation | 0.0244517 | 1 | Phatr_bicluster_0165 |
| GO:0006260 | DNA replication | 0.0624567 | 1 | Phatr_bicluster_0165 |
| GO:0015031 | protein transport | 0.074057 | 1 | Phatr_bicluster_0165 |
| GO:0009765 | photosynthesis light harvesting | 0.0923489 | 1 | Phatr_bicluster_0165 |
| GO:0006508 | proteolysis and peptidolysis | 0.1023 | 1 | Phatr_bicluster_0165 |
| GO:0006457 | protein folding | 0.254387 | 1 | Phatr_bicluster_0165 |
| GO:0006468 | protein amino acid phosphorylation | 0.352461 | 1 | Phatr_bicluster_0165 |
| GO:0006355 | regulation of transcription, DNA-dependent | 0.426896 | 1 | Phatr_bicluster_0165 |
| GO:0006118 | electron transport | 0.441702 | 1 | Phatr_bicluster_0165 |
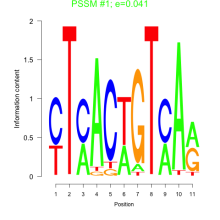


Comments