Phatr_bicluster_0180 Residual: 0.25
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0180 | 0.25 | Phaeodactylum tricornutum |
Displaying 1 - 26 of 26
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12172 riboflavin kinase (Flavokinase)
PHATRDRAFT_12330 2c-methyl-d-erythritol-cyclodiphosphate synthase (MECDP_synthase)
PHATRDRAFT_16962 ribosomal subunit interface protein (RaiA)
PHATRDRAFT_18335 insulin-degrading enzyme (Ptr)
PHATRDRAFT_20124 small gtp-binding protein (Obg_CgtA)
PHATRDRAFT_23552 nicotinamide nucleotide transhydrogenase (pntA)
PHATRDRAFT_29702 ure urease (PLN02303)
PHATRDRAFT_35977 cog4399: uncharacterized protein conserved in bacteria (COG4399 superfamily)
PHATRDRAFT_36048 vde_spiolviolaxanthin de-chloroplast precursor (VDE superfamily)
PHATRDRAFT_38067 anaerobic glycerol-3-phosphate dehydrogenase protein (COG0579)
PHATRDRAFT_40048 ribosome recycling factor (RRF)
PHATRDRAFT_40461 phospholipid glycerol acyltransferase (LPLAT_AGPAT-like)
PHATRDRAFT_41282 ribosome releasing factor (RRF)
PHATRDRAFT_4286 bass family transporter: sodium ion bile acid (COG0385)
PHATRDRAFT_43164 short-chain dehydrogenase reductasesuperfamily protein (NADB_Rossmann superfamily)
PHATRDRAFT_45515 (WcaG)
PHATRDRAFT_46097 chloride channel 7 (Voltage_gated_ClC superfamily)
PHATRDRAFT_47513 (2OG-FeII_Oxy_3 superfamily)
PHATRDRAFT_49708 acyltransferase ws dgat mgat family protein (acyl_WS_DGAT)
PHATRDRAFT_5145 urease accessory protein (Fer4_NifH superfamily)
PHATRDRAFT_51608 (Aldo_ket_red)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0016114 | terpenoid biosynthesis | 0.00690418 | 1 | Phatr_bicluster_0180 |
| GO:0006461 | protein complex assembly | 0.0137641 | 1 | Phatr_bicluster_0180 |
| GO:0009231 | riboflavin biosynthesis | 0.02058 | 1 | Phatr_bicluster_0180 |
| GO:0006821 | chloride transport | 0.0239715 | 1 | Phatr_bicluster_0180 |
| GO:0006814 | sodium ion transport | 0.0374288 | 1 | Phatr_bicluster_0180 |
| GO:0015992 | proton transport | 0.0474084 | 1 | Phatr_bicluster_0180 |
| GO:0006807 | nitrogen compound metabolism | 0.0474084 | 1 | Phatr_bicluster_0180 |
| GO:0006412 | protein biosynthesis | 0.104611 | 1 | Phatr_bicluster_0180 |
| GO:0019538 | protein metabolism | 0.117709 | 1 | Phatr_bicluster_0180 |
| GO:0008152 | metabolism | 0.270353 | 1 | Phatr_bicluster_0180 |

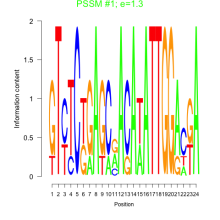
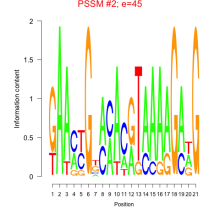
Comments