Phatr_bicluster_0241 Residual: 0.23
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0241 | 0.23 | Phaeodactylum tricornutum |
Displaying 1 - 24 of 24
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_12520 glucose transporter type 3 (Sugar_tr)
PHATRDRAFT_13549 pq-loop repeat family protein transmembrane family protein
PHATRDRAFT_14636 light induced protein like (MPC superfamily)
PHATRDRAFT_15423 dihydroorotate dehydrogenase (DHOD_2_like)
PHATRDRAFT_15844 cg17332-isoform a isoform 2 (V-ATPase_H_C superfamily)
PHATRDRAFT_16960 mgc80429 protein (FKBP_C)
PHATRDRAFT_19949 mercuric reductase (PRK06370)
PHATRDRAFT_27719 at5g60160 f15l12_20 (M18_DAP)
PHATRDRAFT_42688 mitochondrial protein (PPR_2)
PHATRDRAFT_42723 pyrimidine 5-nucleotidase (HAD_2)
PHATRDRAFT_42749 component of oligomeric golgi complex 5 isoform 3
PHATRDRAFT_43076 hd phosphohydrolase family protein (HDc superfamily)
PHATRDRAFT_43174 gdp-fucose transporter (VRG4)
PHATRDRAFT_44816 (DUF938 superfamily)
PHATRDRAFT_46780 metal cationzinc (zn2+)-iron (fe2+) permeasefamily (Zip)
PHATRDRAFT_48865 alpha-l-fucosidase precursor (Alpha_L_fucos)
PHATRDRAFT_49185 transmembrane and coiled-coil domains (DUF106 superfamily)
PHATRDRAFT_51869 inositol 2-dehydrogenase (MviM)
PHATRDRAFT_9267 fkbp2_yarli fk506-binding protein 2 precursor (peptidyl-prolyl cis-trans isomerase) (FKBP_C)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006207 | 'de novo' pyrimidine base biosynthesis | 0.00984608 | 1 | Phatr_bicluster_0241 |
| GO:0006457 | protein folding | 0.0320145 | 1 | Phatr_bicluster_0241 |
| GO:0030001 | metal ion transport | 0.0388631 | 1 | Phatr_bicluster_0241 |
| GO:0008643 | carbohydrate transport | 0.0388631 | 1 | Phatr_bicluster_0241 |
| GO:0005975 | carbohydrate metabolism | 0.14529 | 1 | Phatr_bicluster_0241 |
| GO:0006810 | transport | 0.38354 | 1 | Phatr_bicluster_0241 |
| GO:0006508 | proteolysis and peptidolysis | 0.434343 | 1 | Phatr_bicluster_0241 |
| GO:0006118 | electron transport | 0.441702 | 1 | Phatr_bicluster_0241 |
| GO:0008152 | metabolism | 0.529843 | 1 | Phatr_bicluster_0241 |

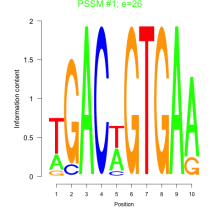

Comments