Regulatory Modules
Displaying 141 - 150 of 170Desulfovibrio vulgaris Hildenborough
| Title | Motif 1 | Motif 2 | Residual | Bicluster Genes | Selected GO Terms |
|---|---|---|---|---|---|
| bicluster_0201 |
1900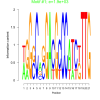 |
10000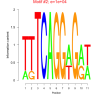 |
0.50 | DVU0655, DVU1437, DVU3213, DVUA0097, DVU0069, DVU0256, DVU0396, DVU0803, DVU0963, DVU1030, DVU1410, DVU1438, DVU2005, DVU2547, DVU2549, DVU2550, DVU2894, DVU3337 | cytoplasm, binding, organic cyclic compound binding, heterocyclic compound binding |
| bicluster_0164 |
2300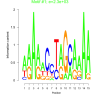 |
4000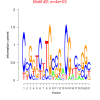 |
0.38 | DVU0074, DVU0657, DVU2227, DVU2323, DVU3174, DVU0153, DVU0157, DVU0321, DVU0792, DVU0793, DVU1041, DVU1242, DVU1407, DVU3186 | TAT protein transport complex, protein complex, racemase and epimerase activity, racemase and epimerase activity, acting ..., cyclic nucleotide binding |
| bicluster_0338 |
2400 |
1800 |
0.39 | DVU1053, DVU1488, DVU1494, DVU1495, DVU1496, DVU1497, DVU1498, DVU1499, DVU1500, DVU1501, DVU1514, DVU1522, DVU1523, DVU1524 | |
| bicluster_0158 |
2700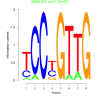 |
2900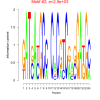 |
0.38 | DVU0857, DVU0865, DVU0994, DVU1267, DVU1602, DVU2470, DVU0242, DVU0758, DVU1278, DVU1336, DVU1337, DVU1874, DVU2310 | ATP binding, adenyl ribonucleotide binding, adenyl nucleotide binding, purine ribonucleoside triphosphate bindi..., nucleoside binding |
| bicluster_0058 |
2700 |
4800 |
0.38 | DVU0197, DVU0206, DVU0207, DVU0209, DVU0210, DVU0211, DVU0212, DVU0214, DVU0216, DVU0217, DVU0219, DVU2850, DVU2851, DVU2853, DVU2857, DVU2858, DVU2859, DVU2861, DVU2864, DVU2860 | |
| bicluster_0287 |
2900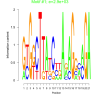 |
13000 |
0.31 | DVU0419, DVU0572, DVU0813, DVU1457, DVU1601, DVU1875, DVU1876, DVU1984, DVU2325, DVU2497, DVU2893, DVU2975 | protein metabolic process, cellular protein metabolic process, macromolecule metabolic process, macromolecule catabolic process, copper ion binding |
| bicluster_0120 |
2900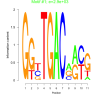 |
16000 |
0.49 | DVU0370, DVU0758, DVU1278, DVU1336, DVU3346, DVU0298, DVU0421, DVU0526, DVU0590, DVU0662, DVU0664, DVU1212, DVU1213, DVU2555, DVU3345 | O-acyltransferase activity, antiporter activity, acetyltransferase activity, secondary active transmembrane transport..., hydrolase activity, acting on carbon-nit... |
| bicluster_0001 |
3100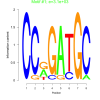 |
11000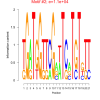 |
0.41 | DVU0383, DVU0824, DVU0964, DVU1243, DVU1790, DVU1962, DVU2120, DVU2139, DVU2169, DVU2326, DVU2602, DVU2622, DVU2915 | glutamate metabolic process, dicarboxylic acid metabolic process, cellular amino acid catabolic process, organic acid catabolic process, small molecule catabolic process |
| bicluster_0015 |
3200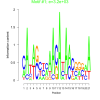 |
17000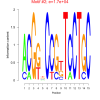 |
0.50 | DVU0022, DVU0117, DVU0286, DVU0539, DVU0621, DVU0681, DVU0807, DVU1163, DVU2023, DVU2041, DVU2502, DVU2661, DVU2736, DVU2906, DVU2994, DVU3000, DVU3007, DVU3128, DVUA0125 | DNA repair, cellular response to DNA damage stimulus, cellular response to stress, DNA metabolic process, copper ion binding |
| bicluster_0193 |
3700 |
600 |
0.46 | DVU0069, DVU0488, DVU0668, DVU2217, DVU2549, DVU2550, DVU3082, DVU0316, DVU0494, DVU0514, DVU1602, DVU1940, DVU1941, DVU1957, DVU2326, DVU3162, DVU3257, DVU3387, DVUA0092 | cofactor binding, N-acetyltransferase activity, N-acyltransferase activity, acetyltransferase activity, pyridoxal phosphate binding |

