|
bicluster_0201 |
1900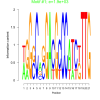 |
10000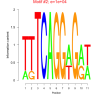 |
0.50 |
DVU0655, DVU1437, DVU3213, DVUA0097, DVU0069, DVU0256, DVU0396, DVU0803, DVU0963, DVU1030, DVU1410, DVU1438, DVU2005, DVU2547, DVU2549, DVU2550, DVU2894, DVU3337 |
cytoplasm, binding, organic cyclic compound binding, heterocyclic compound binding |
|
bicluster_0146 |
0.051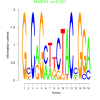 |
81000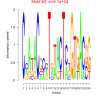 |
0.50 |
DVU0841, DVU0899, DVU0936, DVU0965, DVU0966, DVU1038, DVU1067, DVU1238, DVU1392, DVU1615, DVU1910, DVU1990, DVU2150, DVU2364, DVU2783, DVU2940, DVU3171, DVU3187, DVU3238, DVU3277, DVU0041, DVU0896, DVU1064, DVU1406, DVU1423, DVU1954, DVU2056, DVU2251, DVU2782, DVU2791 |
intramolecular oxidoreductase activity, intramolecular oxidoreductase activity, ..., NADP binding |
|
bicluster_0014 |
150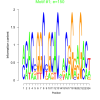 |
1300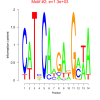 |
0.50 |
DVU0072, DVU0076, DVU0080, DVU0328, DVU0425, DVU0428, DVU0439, DVU0441, DVU0478, DVU0638, DVU0680, DVU0691, DVU0743, DVU1004, DVU1051, DVU1064, DVU1423, DVU1425, DVU1452, DVU1607, DVU1926, DVU2568, DVU2583, DVU2627, DVU2673, DVU2779, DVU3113, DVU3127, DVU3296 |
organonitrogen compound catabolic proces..., single-organism catabolic process, cellular catabolic process, catabolic process, organic substance catabolic process |
|
bicluster_0015 |
3200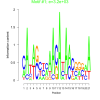 |
17000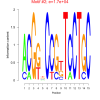 |
0.50 |
DVU0022, DVU0117, DVU0286, DVU0539, DVU0621, DVU0681, DVU0807, DVU1163, DVU2023, DVU2041, DVU2502, DVU2661, DVU2736, DVU2906, DVU2994, DVU3000, DVU3007, DVU3128, DVUA0125 |
DNA repair, cellular response to DNA damage stimulus, cellular response to stress, DNA metabolic process, copper ion binding |
|
mex_bicluster_0279 |
340 |
6200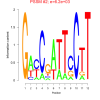 |
0.50 |
6828169, 6830144, 6826378, 6829190, 6831000, RMQ12506, 6826419, 6826294, 6828410, 6827787, 6827517, 6827830, 6829552, 6827786, 6827986, 6826400, 6829831, 6827814, 6827041, RMQ09372, 6827907, 6827521, 6829460, 6827785, 6829848 |
|
|
bicluster_0244 |
0.0000000065 |
0.0051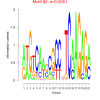 |
0.50 |
DVU2198, DVU0909, DVU0915, DVU0920, DVU0921, DVU0926, DVU0929, DVU1098, DVU1707, DVU1708, DVU1709, DVU1736, DVU1988, DVU2249, DVU2250, DVU2296, DVU2645, DVU2646, DVU2831 |
cell wall organization, cell wall organization or biogenesis, external encapsulating structure organiz... |
|
bicluster_0284 |
70 |
500 |
0.50 |
DVU0181, DVU0185, DVU0283, DVU1416, DVU2334, DVU2346, DVU0008, DVU0877, DVU1899, DVU2147, DVU2204, DVU2303, DVU3000, DVU3007, DVU3034, DVU3057, DVU3214 |
DNA recombination, DNA metabolic process, cofactor binding, oxidoreductase activity, acting on the C..., pyridoxal phosphate binding |
|
bicluster_0121 |
17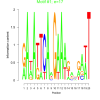 |
2200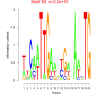 |
0.50 |
DVU0408, DVU1171, DVU1367, DVU1387, DVU1739, DVU1750, DVU1995, DVU2017, DVU2068, DVU2178, DVU2263, DVU2682, DVU2842, DVU3138, DVU3218, DVUA0018, DVUA0067, DVUA0135, DVU0196, DVU0563, DVU0802, DVU0993, DVU1773, DVU2671, DVU3038, DVUA0071 |
DNA integration, nucleic acid metabolic process, TAT protein transport complex, DNA (cytosine-5-)-methyltransferase acti..., DNA-methyltransferase activity |
|
bicluster_0122 |
0.0053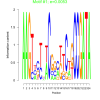 |
42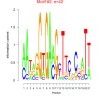 |
0.49 |
DVU0033, DVU0122, DVU0123, DVU0583, DVU0598, DVU0599, DVU0935, DVU0942, DVU1241, DVU1786, DVU1854, DVU1880, DVU2444, DVU2445, DVU2516, DVU0523, DVU0524, DVU0976, DVU0977, DVU1073, DVU1857, DVU1869, DVU2948, DVU2952, DVU3125, DVU3218 |
response to extracellular stimulus, cellular response to extracellular stimu..., cellular response to external stimulus, cellular response to stress, cell communication |
|
bicluster_0053 |
0.29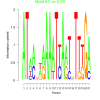 |
44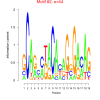 |
0.49 |
DVU0146, DVU0148, DVU0149, DVU0150, DVU0152, DVU0372, DVU0458, DVU0593, DVU0678, DVU0751, DVU0805, DVU0806, DVU0882, DVU1670, DVU2014, DVU2061, DVU2104, DVU2952, DVU3125, DVU3182, DVU3188, DVU3384, DVUA0036, DVUA0061, DVU0147, DVU0594, DVU2088 |
integral component of membrane, intrinsic component of membrane, FMN binding |