Organism : Desulfovibrio vulgaris Hildenborough
| Module List :


DVU0140
response regulator
Functional Annotations (3)
| Function | System |
|---|---|
| two-component response regulator activity | go/ molecular_function |
| two-component signal transduction system (phosphorelay) | go/ biological_process |
| regulation of transcription, DNA-dependent | go/ biological_process |

Regulation information for DVU0140
(Mouseover regulator name to see its description)
| Regulator | Module | Operator |
|---|---|---|
| DVU0679 | 37 | tf |
| DVU0744 | 37 | tf |
| DVU1063 | 37 | tf |
| DVU1340 | 37 | tf |
| DVU1628 | 37 | tf |
| DVU1949 | 37 | tf |
| DVU2359 | 37 | tf |
| DVU2547 DVU2582 |
37 | combiner |
| DVU2547 DVU3229 |
37 | combiner |
| DVU2934 | 37 | tf |
| DVU3111 | 37 | tf |
| DVU3167 | 37 | tf |
| DVU3381 | 37 | tf |
| DVUA0100 | 37 | tf |
| DVU1730 | 99 | tf |
| DVU2423 DVU3381 |
99 | combiner |
| DVU2644 DVU1584 |
99 | combiner |
| DVU3255 DVU1584 |
99 | combiner |
| DVUA0151 DVU1730 |
99 | combiner |

Motif information (de novo identified motifs for modules)
There are 4 motifs predicted.
Click on the RegPredict links to explore the motif in RegPredict.
| Motif Id | e-value | Consensus | Motif Logo | RegPredict |
|---|---|---|---|---|
| 73 | 6.10e+03 | ATtGCTaTCGT |
 |
RegPredict |
| 74 | 1.60e+04 | CAtAtgacAGAaaCC |
 |
RegPredict |
| 191 | 0.00e+00 | aagT.tTcacagctTCtgcCGATa |
 |
RegPredict |
| 192 | 4.00e-02 | ATtgtgAtACtTtaCccATtG |
 |
RegPredict |

Functional Enrichment for DVU0140
| Function | System |
|---|---|
| two-component response regulator activity | go/ molecular_function |
| two-component signal transduction system (phosphorelay) | go/ biological_process |
| regulation of transcription, DNA-dependent | go/ biological_process |

Module neighborhood information for DVU0140
| Gene | Common Name | Description | Module membership |
|---|---|---|---|
| DVU0020 | hypothetical protein DVU0020 | 99, 314 | |
| DVU0040 | hypothetical protein DVU0040 | 37, 267 | |
| DVU0140 | response regulator | 37, 99 | |
| DVU0145 | response regulator | 37, 205 | |
| DVU0147 | lipoprotein | 37, 53 | |
| DVU0148 | lipoprotein | 53, 99 | |
| DVU0149 | hypothetical protein DVU0149 | 53, 99 | |
| DVU0150 | hypothetical protein DVU0150 | 53, 99 | |
| DVU0151 | HAMP domain/sigma-54 interaction domain-containing protein | 17, 99 | |
| DVU0152 | phosphoenolpyruvate synthase-like protein | 53, 99 | |
| DVU0218 | tail protein | 37, 314 | |
| DVU0231 | hypothetical protein DVU0231 | 99, 106 | |
| DVU0232 | hypothetical protein DVU0232 | 37, 99 | |
| DVU0257 | acetyltransferase | 37, 69 | |
| DVU0301 | hypothetical protein DVU0301 | 37, 330 | |
| DVU0358 | hypothetical protein DVU0358 | 37, 99 | |
| DVU0360 | ilvB-1 | acetolactate synthase catalytic subunit | 99, 247 |
| DVU0361 | acetolactate synthase 1 regulatory subunit | 99, 205 | |
| DVU0381 | nhaC-1 | Na+/H+ antiporter NhaC | 37, 341 |
| DVU0391 | hypothetical protein DVU0391 | 99, 161 | |
| DVU0392 | aromatic aminotransferase | 37, 185 | |
| DVU0394 | radical SAM domain-containing protein | 62, 99 | |
| DVU0425 | hypothetical protein DVU0425 | 14, 37 | |
| DVU0638 | hypothetical protein DVU0638 | 14, 37 | |
| DVU0639 | pomB | chemotaxis protein PomB | 37, 205 |
| DVU0641 | hypothetical protein DVU0641 | 37, 225 | |
| DVU0644 | hypothetical protein DVU0644 | 99, 222 | |
| DVU0672 | hypothetical protein DVU0672 | 99, 230 | |
| DVU0976 | response regulator | 99, 122 | |
| DVU0977 | hypothetical protein DVU0977 | 99, 122 | |
| DVU0989 | periplasmic divalent cation tolerance protein cutA | 57, 99 | |
| DVU1011 | hypothetical protein DVU1011 | 37, 245 | |
| DVU1072 | hypothetical protein DVU1072 | 99, 236 | |
| DVU1073 | hypothetical protein DVU1073 | 99, 122 | |
| DVU1126 | lipoprotein | 37, 267 | |
| DVU1259 | hypothetical protein DVU1259 | 46, 99 | |
| DVU1360 | galE | UDP-glucose 4-epimerase | 37, 342 |
| DVU1442 | flagellin FlaG | 37, 286 | |
| DVU1604 | hypothetical protein DVU1604 | 37, 181 | |
| DVU1691 | hypothetical protein DVU1691 | 37, 44 | |
| DVU1824 | hypothetical protein DVU1824 | 27, 99 | |
| DVU1906 | hypothetical protein DVU1906 | 37, 313 | |
| DVU1993 | cation transporter E1-E2 family ATPase | 37, 296 | |
| DVU2095 | thiS | thiamine biosynthesis protein ThiS | 99, 345 |
| DVU2100 | universal stress protein | 66, 99 | |
| DVU2267 | hypothetical protein DVU2267 | 37, 179 | |
| DVU2311 | None | 37, 261 | |
| DVU2473 | hypothetical protein DVU2473 | 37, 46 | |
| DVU2498 | hypothetical protein DVU2498 | 37, 342 | |
| DVU2551 | HD domain-containing protein | 37, 342 | |
| DVU2589 | hypothetical protein DVU2589 | 37, 153 | |
| DVU2590 | sensory box protein | 37, 153 | |
| DVU2666 | phosphate ABC transporter permease | 99, 214 | |
| DVU2972 | chemotaxis protein CheD | 37, 274 | |
| DVU3021 | HDIG domain-containing protein | 37, 245 | |
| DVU3036 | hypothetical protein DVU3036 | 37, 281 | |
| DVU3060 | hypothetical protein DVU3060 | 37, 166 | |
| DVU3073 | hypothetical protein DVU3073 | 99, 211 | |
| DVU3111 | Crp/FNR family transcriptional regulator | 37, 267 | |
| DVU3225 | hypothetical protein DVU3225 | 37, 198 | |
| DVU3342 | hypothetical protein DVU3342 | 37, 123 | |
| DVU3391 | hypothetical protein DVU3391 | 37, 305 |
Gene Page Help
Network Tab
If the gene is associated with a module(s), its connection to given modules along with other members of that module are shown as network by using CytoscapeWeb. In this view, each green colored circular nodes represent module member genes, purple colored diamonds represent module motifs and red triangles represent regulators. Each node is connected to module (Bicluster) via edges. This representation provides quick overview of all genes, regulators and motifs for modules. It also allows one to see shared genes/motifs/regulators among diferent modules.
Network representation is interactive. You can zoom in/out and move nodes/edges around. Clicking on a node will open up a window to give more details. For genes, Locus tag, organism, genomic coordinates, NCBI gene ID, whether it is transcription factor or not and any associated functional information will be shown. For regulators, number of modules are shown in addition to gene details. For motifs, e-value, consensus sequence and sequence logo will be shown. For modules, expression profile plot, motif information, functional associations and motif locations for each member of the module will be shown.
You can pin information boxes by using button in the box title and open up additional ones on the same screen for comparative analysis.
Regulation Tab
Regulation tab for each gene includes regulatory influences such as environmental factors or transcription factors or their combinations identified by regulatory network inference algorithms.
If the gene is a member of a module, regulators influencing that module are also considered to regulate the gene. Regulators table list total number of regulatory influences, regulators, modules and type of the influence.
You can see description of the regulator inside the tooltip when you mouseover. In certain cases the regulatory influence is predicted to be the result of the combination of two influences. These are indicated as combiner in the column labeled "Operator".
For transcription factors, an additional table next to regulator table will be show. This table show modules that are influenced by the transcription factor.
Motifs Tab
Network inference algorithm uses de novo motif prediction for assigning genes to modules. If there are any motifs identified in the upstream region of a gene, the motif will be shown here. For each motif sequence logo, consensus and e-value will be shown.
Functions Tab
Identification of functional enrichment for the module members is important in associating predicted motifs and regulatory influences with pathways. As described above, the network inference pipeline includes a functional enrichment module by which hypergeometric p-values are used to identify over representation of functional ontology terms among module members.
Network Portal presents functional ontologies from KEGG, GO, TIGRFAM, and COG as separate tables that include function name, type, corrected and uncorrected hypergeometric p-values, and the number of genes assigned to this category out of total number of genes in the module.
Module Members Tab
Identity of gene members in a module may help to identify potential interactions between different functional modules. Therefore, neighbor genes that share the same module(s) with gene under consideration are shown here. For each memebr, gene name, description and modules that contain it are listed.
Help Tab
This help page. More general help can be accessed by clicking help menu in the main navigation bar.
CircVis
Our circular module explorer is adapted from visquick originally developed by Dick Kreisberg of Ilya Shmulevich lab at ISB for The Cancer Genome Atlas. We use simplified version of visquick to display distribution of module members and their interactions across the genome. This view provides summary of regulation information for a gene. The main components are;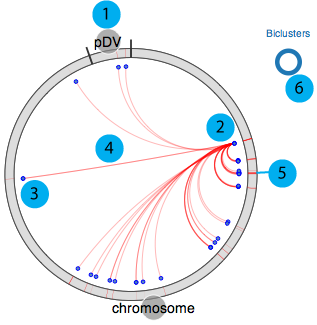
- 1. All genomic elements for the organism are represented as a circle and each element is separated by black tick marks. In this example chromosome and pDV represent main chromosome and plasmid for D. vulgaris Hildenborough, respectively.
- 2. Source gene
- 3. Target genes (other module members)
- 4. Interactions between source and target genes for a particular module
- 5. Module(s) that source gene and target genes belong to
- 6. Visualisation legend


 Module member
Module member  Regulator
Regulator  Motif
Motif 
Social Tab
Network Portal is designed to promote collaboration through social interactions. Therefore interested researchers can share information, questions and updates for a particular gene.
Users can use their Disqus, Facebook, Twitter or Google accounts to connect to this page (We recommend Google). Each module and gene page includes comments tab that lists history of the interactions for that gene. You can browse the history, make updates, raise questions and share these activities with social web.
In the next releases of the network portal, we are planning to create personal space for each user where you can share you space that contains all the analysis steps you did along with relevant information.