Regulatory Modules
Displaying 161 - 170 of 170Desulfovibrio vulgaris Hildenborough
| Title | Motif 1 | Motif 2 | Residual | Bicluster Genes | Selected GO Terms |
|---|---|---|---|---|---|
| bicluster_0263 |
0.00000016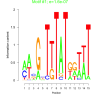 |
0.036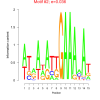 |
0.49 | DVU0030, DVU1564, DVU2452, DVU2745, DVU0369, DVU0560, DVU0618, DVU1257, DVU1561, DVU1690, DVU1705, DVU2000, DVU2007, DVU2096, DVU2106, DVU2182, DVU2183, DVU2198, DVU2451, DVU2651, DVU2718, DVU2778, DVUA0028 | integral component of plasma membrane, intrinsic component of plasma membrane, plasma membrane part, nucleic acid binding, DNA binding |
| bicluster_0232 |
0.04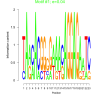 |
3.5 |
0.49 | DVU0302, DVU1190, DVU1372, DVU1373, DVU1374, DVU1375, DVU1377, DVU1378, DVU2532, DVU2965, DVU0837, DVU0838, DVU0936, DVU0942, DVU0986, DVU0987, DVU1093, DVU1367, DVU1392, DVU1612, DVU1834, DVU3187 | RNA processing, ncRNA processing, ncRNA metabolic process, cell cycle, TAT protein transport complex |
| bicluster_0201 |
1900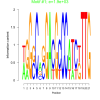 |
10000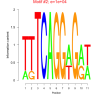 |
0.50 | DVU0655, DVU1437, DVU3213, DVUA0097, DVU0069, DVU0256, DVU0396, DVU0803, DVU0963, DVU1030, DVU1410, DVU1438, DVU2005, DVU2547, DVU2549, DVU2550, DVU2894, DVU3337 | cytoplasm, binding, organic cyclic compound binding, heterocyclic compound binding |
| bicluster_0146 |
0.051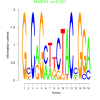 |
81000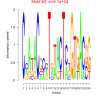 |
0.50 | DVU0841, DVU0899, DVU0936, DVU0965, DVU0966, DVU1038, DVU1067, DVU1238, DVU1392, DVU1615, DVU1910, DVU1990, DVU2150, DVU2364, DVU2783, DVU2940, DVU3171, DVU3187, DVU3238, DVU3277, DVU0041, DVU0896, DVU1064, DVU1406, DVU1423, DVU1954, DVU2056, DVU2251, DVU2782, DVU2791 | intramolecular oxidoreductase activity, intramolecular oxidoreductase activity, ..., NADP binding |
| bicluster_0014 |
150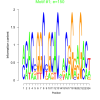 |
1300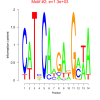 |
0.50 | DVU0072, DVU0076, DVU0080, DVU0328, DVU0425, DVU0428, DVU0439, DVU0441, DVU0478, DVU0638, DVU0680, DVU0691, DVU0743, DVU1004, DVU1051, DVU1064, DVU1423, DVU1425, DVU1452, DVU1607, DVU1926, DVU2568, DVU2583, DVU2627, DVU2673, DVU2779, DVU3113, DVU3127, DVU3296 | organonitrogen compound catabolic proces..., single-organism catabolic process, cellular catabolic process, catabolic process, organic substance catabolic process |
| bicluster_0015 |
3200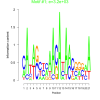 |
17000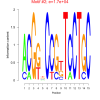 |
0.50 | DVU0022, DVU0117, DVU0286, DVU0539, DVU0621, DVU0681, DVU0807, DVU1163, DVU2023, DVU2041, DVU2502, DVU2661, DVU2736, DVU2906, DVU2994, DVU3000, DVU3007, DVU3128, DVUA0125 | DNA repair, cellular response to DNA damage stimulus, cellular response to stress, DNA metabolic process, copper ion binding |
| bicluster_0244 |
0.0000000065 |
0.0051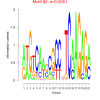 |
0.50 | DVU2198, DVU0909, DVU0915, DVU0920, DVU0921, DVU0926, DVU0929, DVU1098, DVU1707, DVU1708, DVU1709, DVU1736, DVU1988, DVU2249, DVU2250, DVU2296, DVU2645, DVU2646, DVU2831 | cell wall organization, cell wall organization or biogenesis, external encapsulating structure organiz... |
| bicluster_0284 |
70 |
500 |
0.50 | DVU0181, DVU0185, DVU0283, DVU1416, DVU2334, DVU2346, DVU0008, DVU0877, DVU1899, DVU2147, DVU2204, DVU2303, DVU3000, DVU3007, DVU3034, DVU3057, DVU3214 | DNA recombination, DNA metabolic process, cofactor binding, oxidoreductase activity, acting on the C..., pyridoxal phosphate binding |
| bicluster_0121 |
17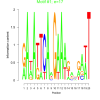 |
2200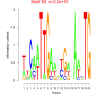 |
0.50 | DVU0408, DVU1171, DVU1367, DVU1387, DVU1739, DVU1750, DVU1995, DVU2017, DVU2068, DVU2178, DVU2263, DVU2682, DVU2842, DVU3138, DVU3218, DVUA0018, DVUA0067, DVUA0135, DVU0196, DVU0563, DVU0802, DVU0993, DVU1773, DVU2671, DVU3038, DVUA0071 | DNA integration, nucleic acid metabolic process, TAT protein transport complex, DNA (cytosine-5-)-methyltransferase acti..., DNA-methyltransferase activity |
| bicluster_0029 |
0.67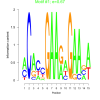 |
32000 |
0.50 | DVU0002, DVU0003, DVU0237, DVU0279, DVU0502, DVU0809, DVU0810, DVU1195, DVU1248, DVU1576, DVU1584, DVU1661, DVU1666, DVU1889, DVU1893, DVU2339, DVU3275, DVU3307, DVU3395, DVU0004, DVU0161, DVU0162, DVU1062 | carbohydrate derivative biosynthetic pro..., organophosphate biosynthetic process, carbohydrate derivative metabolic proces..., phosphate-containing compound metabolic ..., organophosphate metabolic process |

