Phatr_bicluster_0179 Residual: 0.41
Phaeodactylum tricornutum
| Title | Residual | Model version | Organism |
|---|---|---|---|
| Phatr_bicluster_0179 | 0.41 | Phaeodactylum tricornutum |
Displaying 1 - 27 of 27
" class="views-fluidgrid-wrapper clear-block">
PHATRDRAFT_10005 phosphopantothenoylcysteine synthetase (DFP superfamily)
PHATRDRAFT_28568 snf2 domain-containing protein helicase domain-containing protein ring finger domain-containing protein (SNF2_N)
PHATRDRAFT_33186 tsr2 protein (WGG superfamily)
PHATRDRAFT_33584 tyw23_orysj trna wybutosine-synthesizing protein 2 3 4 (TYW3 superfamily)
PHATRDRAFT_40691 nitrate transporter
PHATRDRAFT_41259 aaa familycdc48 subfamily (AAA)
PHATRDRAFT_41639 retinitis pigmentosa gtpase regulator
PHATRDRAFT_42963 ac006341_12 ests gb (DUF2358 superfamily)
PHATRDRAFT_44876 generic methyltransferase (ovoA_Nterm)
PHATRDRAFT_45108 (COG1258)
PHATRDRAFT_45192 adenylate kinase (adk)
PHATRDRAFT_45251 pseudouridine synthase (PseudoU_synth_ScPUS7)
PHATRDRAFT_46436 cellulose-family ii
PHATRDRAFT_46703 (Senescence)
PHATRDRAFT_47358 plastid-lipid associated protein pap (PAP_fibrillin superfamily)
PHATRDRAFT_51092 glutamine synthetase (PLN02284)
PHATRDRAFT_54681 glycosidefamily 16 (GH16_laminarinase_like)
PHATRDRAFT_54983 nitrate reductase (PLN02252)
PHATRDRAFT_6457 cytidylate kinase (UMP_CMP_kin_fam)
PHATRDRAFT_7893 ribosomal protein s15 (Ribosomal_S15p_S13e)
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006139 | nucleobase, nucleoside, nucleotide and nucleic acid metabolism | 0.00018625 | 0.427444 | Phatr_bicluster_0179 |
| GO:0006542 | glutamine biosynthesis | 0.0019758 | 1 | Phatr_bicluster_0179 |
| GO:0006807 | nitrogen compound metabolism | 0.0273521 | 1 | Phatr_bicluster_0179 |
| GO:0008033 | tRNA processing | 0.0388725 | 1 | Phatr_bicluster_0179 |
| GO:0016567 | protein ubiquitination | 0.13382 | 1 | Phatr_bicluster_0179 |
| GO:0006412 | protein biosynthesis | 0.277391 | 1 | Phatr_bicluster_0179 |
| GO:0006468 | protein amino acid phosphorylation | 0.2936 | 1 | Phatr_bicluster_0179 |

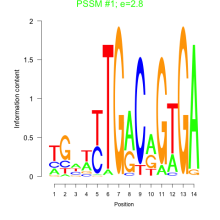
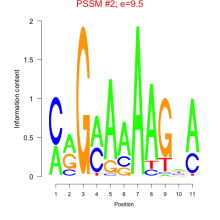
Comments