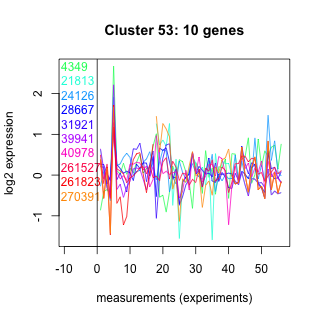
Thaps_hclust_0053 Hierarchical Clustering
Thalassiosira pseudonana
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0009081 | branched chain family amino acid metabolism | 0.00165341 | 1 | Thaps_hclust_0053 |
| GO:0006260 | DNA replication | 0.0286397 | 1 | Thaps_hclust_0053 |
| GO:0005975 | carbohydrate metabolism | 0.0606126 | 1 | Thaps_hclust_0053 |
| GO:0008152 | metabolism | 0.296505 | 1 | Thaps_hclust_0053 |
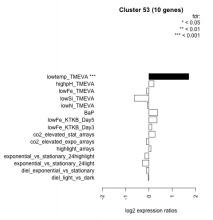
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| diel_light_vs_dark | diel_light_vs_dark | -0.028 | 0.959 |
| lowFe_KTKB_Day3 | lowFe_KTKB_Day3 | 0.125 | 0.75 |
| lowFe_KTKB_Day5 | lowFe_KTKB_Day5 | 0.346 | 0.239 |
| BaP | BaP | 0.377 | 0.227 |
| exponential_vs_stationary_24highlight | exponential_vs_stationary_24highlight | -0.171 | 0.256 |
| co2_elevated_stat_arrays | co2_elevated_stat_arrays | 0.267 | 0.345 |
| lowtemp_TMEVA | lowtemp_TMEVA | 1.710 | 0.000735 |
| highpH_TMEVA | highpH_TMEVA | 0.218 | 0.325 |
| co2_elevated_expo_arrays | co2_elevated_expo_arrays | -0.115 | 0.726 |
| lowFe_TMEVA | lowFe_TMEVA | -0.108 | 0.816 |
| exponential_vs_stationary_24light | exponential_vs_stationary_24light | -0.275 | 0.545 |
| lowN_TMEVA | lowN_TMEVA | -0.065 | 0.866 |
| diel_exponential_vs_stationary | diel_exponential_vs_stationary | -0.109 | 0.707 |
| lowSi_TMEVA | lowSi_TMEVA | -0.630 | 0.298 |
| highlight_arrays | highlight_arrays | 0.089 | 0.72 |