Regulatory Motifs cMonkey Network
| Title | Logo | e.value | Number of Sites | Width | Motif Bicluster | |
|---|---|---|---|---|---|---|
| motif_0168_1 |
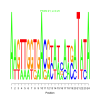 |
0.25 | 4 | 24 | bicluster_0168 | |
| motif_0505_1 |
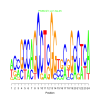 |
0.000016 | 8 | 24 | bicluster_0505 |