Module 6 Residual: 0.61
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0006 | v02 | 0.61 | -5.60 |
| Rv0345 CTP:molybdopterin cytidylyltransferase | Rv0509 Glutamyl-tRNA reductase (EC 1.2.1.70) |
| Rv0510 Porphobilinogen deaminase (EC 2.5.1.61) | Rv0513 Transmembrane transport protein MmpL2 |
| Rv1089 PE family protein | Rv1173 7,8-didemethyl-8-hydroxy-5-deazariboflavin synthase subunit 1 / 7,8-didemethyl-8-hydroxy-5-deazariboflavin synthase subunit 2 |
| Rv1207 Non functional Dihydropteroate synthase 2 | Rv1208 Glycosyltransferases involved in cell wall biogenesis |
| Rv2425c VWA containing CoxE family protein | Rv2477c ABC transporter ATP-binding protein |
| Rv2726c Diaminopimelate epimerase (EC 5.1.1.7) | Rv2995c 3-isopropylmalate dehydrogenase (EC 1.1.1.85) |
| Rv3025c Cysteine desulfurase (EC 2.8.1.7) | Rv3103c |
| Rv3104c Potassium efflux system KefA protein / Small-conductance mechanosensitive channel | Rv3480c Wax ester synthase/acyl-CoA:diacylglycerol acyltransferase |
| Rv3803c Antigen 85-C precursor (85C) (Antigen 85 complex C) (Ag85C) (Mycolyl transferase 85C) (EC 2.3.1.-) | Rv3843c PROBABLE CONSERVED TRANSMEMBRANE PROTEIN |
|
GO:0008883 | glutamyl-tRNA reductase activity |
|
GO:0040007 | growth |
|
GO:0004418 | hydroxymethylbilane synthase activity |
|
GO:0005886 | plasma membrane |
|
GO:0009108 | coenzyme biosynthetic process |
|
GO:0051409 | response to nitrosative stress |
|
GO:0042803 | protein homodimerization activity |
|
GO:0004156 | dihydropteroate synthase activity |
|
GO:0000287 | magnesium ion binding |
|
GO:0006011 | UDP-glucose metabolic process |
|
GO:0016758 | transferase activity, transferring hexosyl groups |
|
GO:0051260 | protein homooligomerization |
|
GO:0070207 | protein homotrimerization |
|
GO:0005829 | cytosol |
|
GO:0008837 | diaminopimelate epimerase activity |
|
GO:0003862 | 3-isopropylmalate dehydrogenase activity |
|
GO:0031071 | cysteine desulfurase activity |
|
GO:0016226 | iron-sulfur cluster assembly |
|
GO:0045017 | glycerolipid biosynthetic process |
|
GO:0047196 | long-chain-alcohol O-fatty-acyltransferase activity |
|
GO:0005576 | extracellular region |




















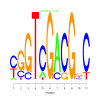
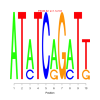
Discussion