Module 117
Summary
| Residual | Gene Count | Condition Count | 1st Q Condition Block * | 5th Q Condition Block * |
|---|---|---|---|---|
| 0.59 | 27 | 63 | In vivo vs. In vitro | NA |
Bicluster expression profile
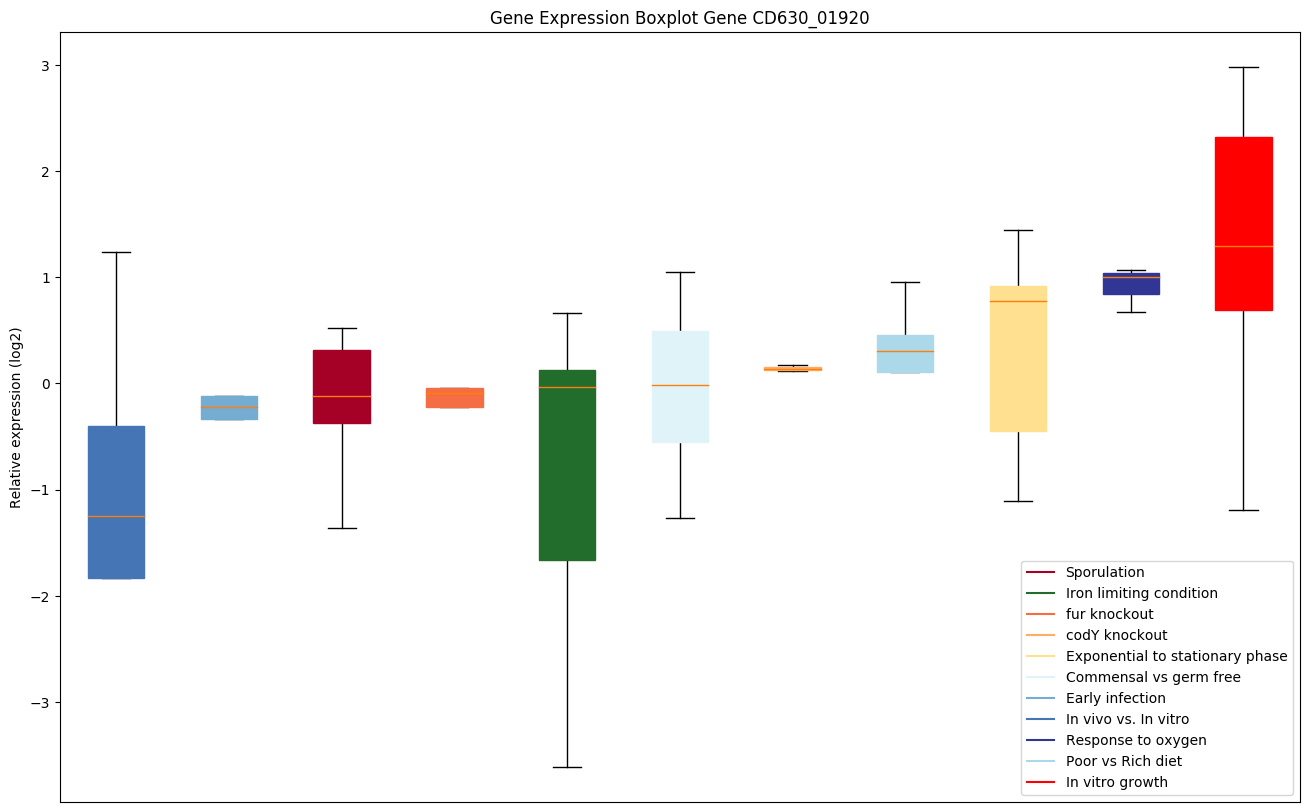
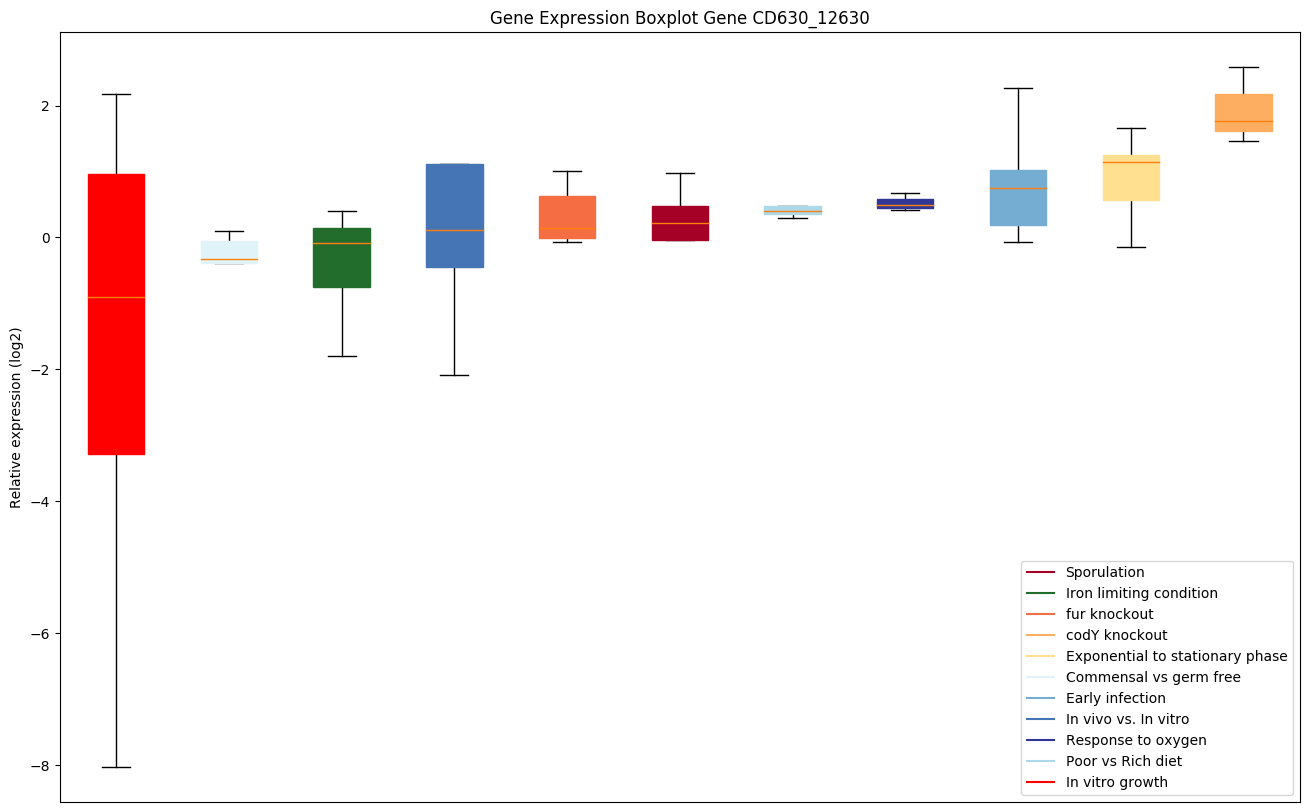
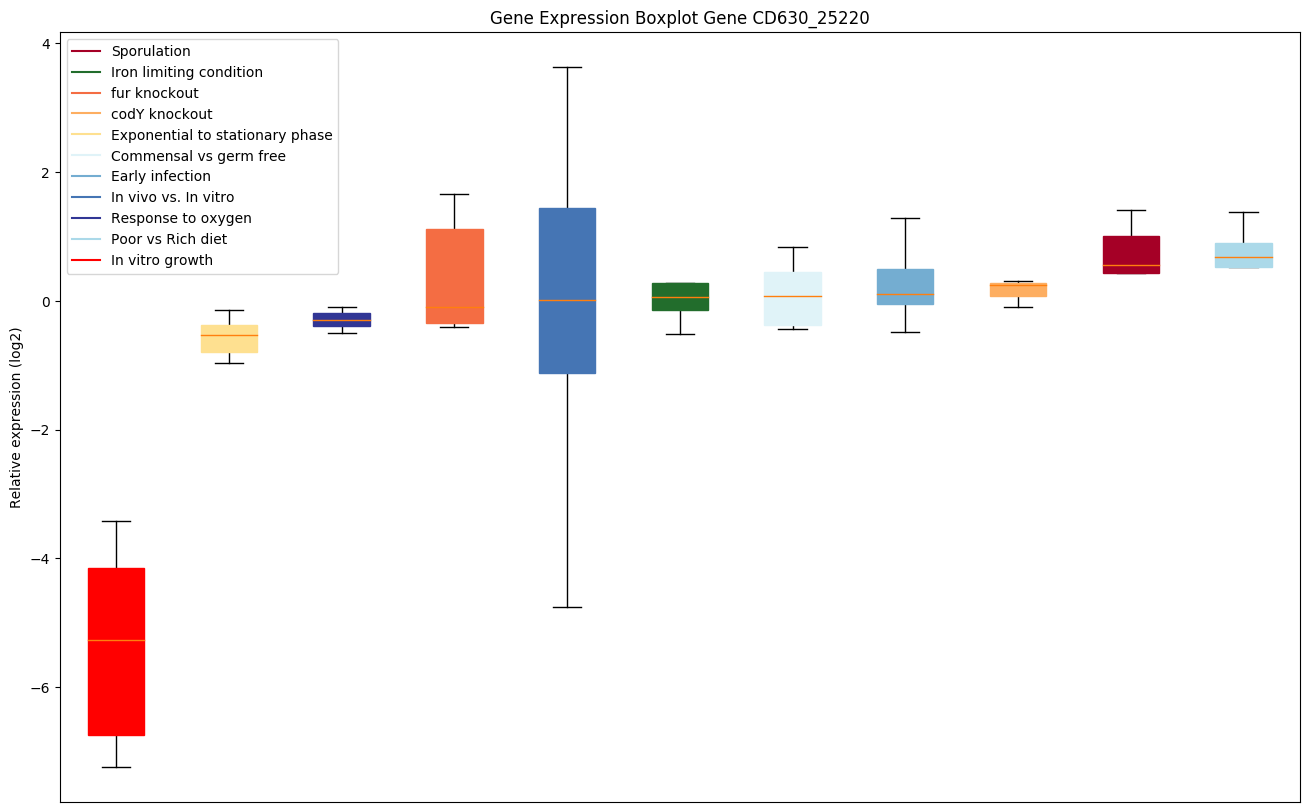
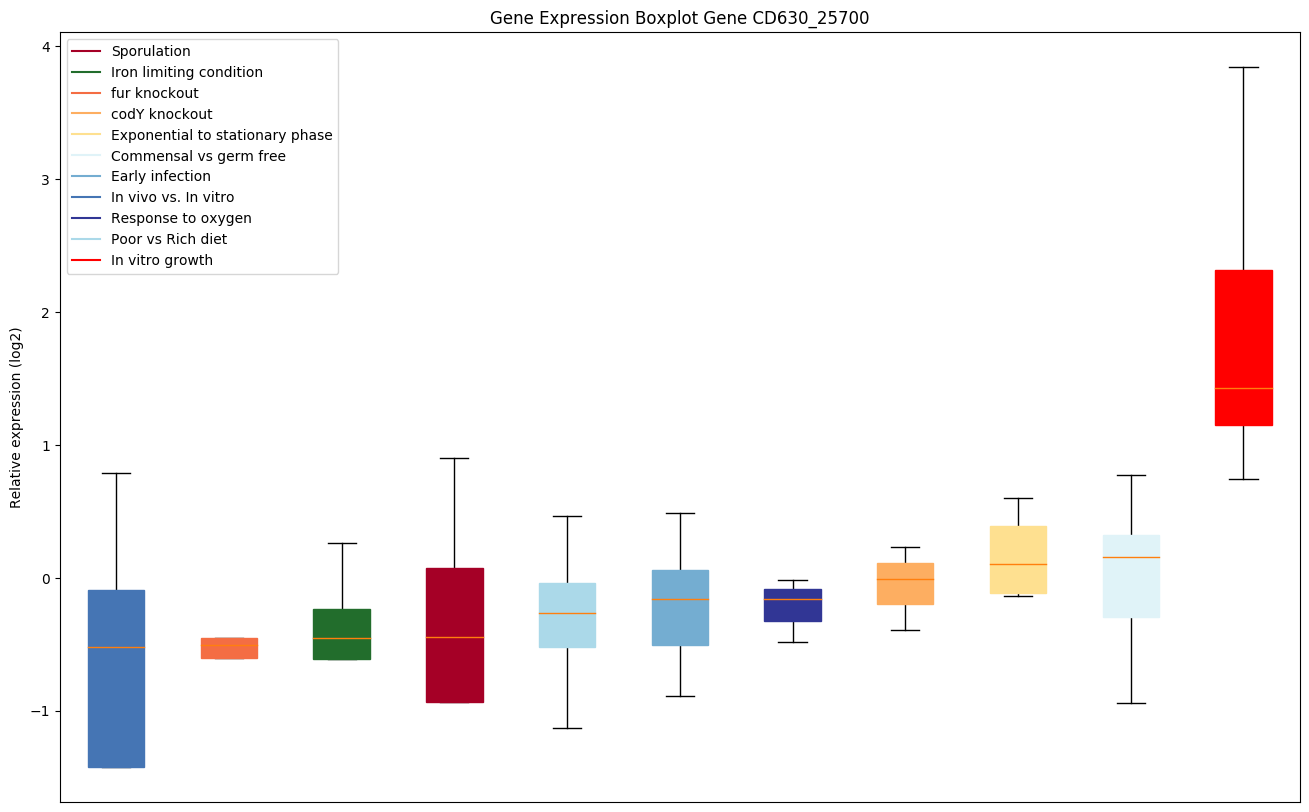
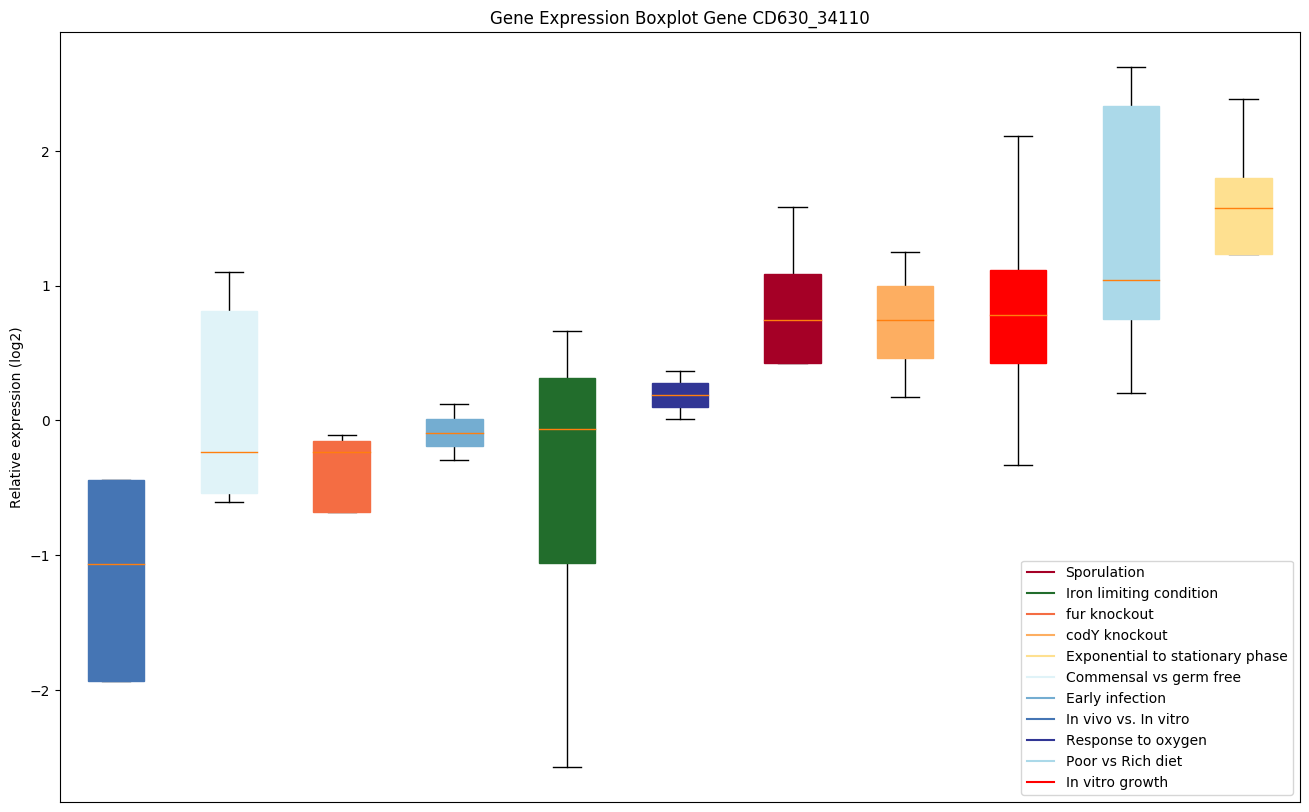
Expression of genes in subset of conditions included in bicluster on left side of red dashed line and out of bicluster on right of red dashed line. Each condition is represented as a boxplot, ordered by their median expression for the bicluster genes (smallest to largest) and colored according to condition blocks.
|
|
de novo identified motifs are listed below
Candidate transcriptional regulators for each module were determined with a linear regression based approach (Inferelator) and by evaluating the statistical significance of the overlap (Hypergeometric) between modules and putative TF and alternative sigma factor regulons (CcpA, CodY, Fur, PrdR, SigB, SigD, SigE, SigF, SigG, SigH, SigK, Spo0A) compiled from available literature.
For filtering high confidence influences, we used hypergeometric test adjusted p-value <=0.05 and minimum four genes in the overlap between the module and the TF regulon. The same thresholds were used for evaluating the functional enrichment.
1. Betas: correspond to the average coefficients of the Bayesian regressions between module expression and TF expression profiles. The values indicate the magnitude and direction (activation or repression for positive or negative values, respectively) of each TF-module interaction.
2. Confidence scores: indicate the likelihood of the TF-module interactions.
The module is significantly enriched with genes associated to the indicated functional terms
Genes that are included in this module
* "Gene essentiality is based on TnSeq Data from: Dembek M, Barquist L, Boinett CJ, et al. High-throughput analysis of gene essentiality and sporulation in Clostridium difficile. mBio. 2015;6(2):e02383. Published 2015 Feb 24.doi:10.1128/mBio.02383-14".
| Title | Short Name | Product | Function | Essentiality * | in vivo Essentiality | Rich broth Essentiality | Expression |
|---|---|---|---|---|---|---|---|
| CD630_01920 | cls | Cardiolipin synthetase 1 | Catalyzes the reversible phosphatidyl group transfer from one phosphatidylglycerol molecule to another to form cardiolipin (CL) (diphosphatidylglycerol) and glycerol. |
 |
|||
| CD630_22870 | Putative transporter |
 |
|||||
| CD630_30440 | Transcriptional regulator, RpiR family |
 |
|||||
| CD630_12630 | Putative FMN-dependent dehydrogenase |
 |
|||||
| CD630_24750 | Putative membrane protein |
 |
|||||
| CD630_12610 | Putative ribonucleotide-diphosphate reductase | Catalyzes the reduction of ribonucleotides to deoxyribonucleotides. May function to provide a pool of deoxyribonucleotide precursors for DNA repair during oxygen limitation and/or for immediate growth after restoration of oxygen. |
 |
||||
| CD630_24770 | Conserved hypothetical protein |
 |
|||||
| CD630_25220 | Conserved hypothetical protein | Functions as a ribosomal silencing factor. Interacts with ribosomal protein L14 (rplN), blocking formation of intersubunit bridge B8. Prevents association of the 30S and 50S ribosomal subunits and the formation of functional ribosomes, thus repressing translation. |
 |
||||
| CD630_10130 | Two-component response regulator |
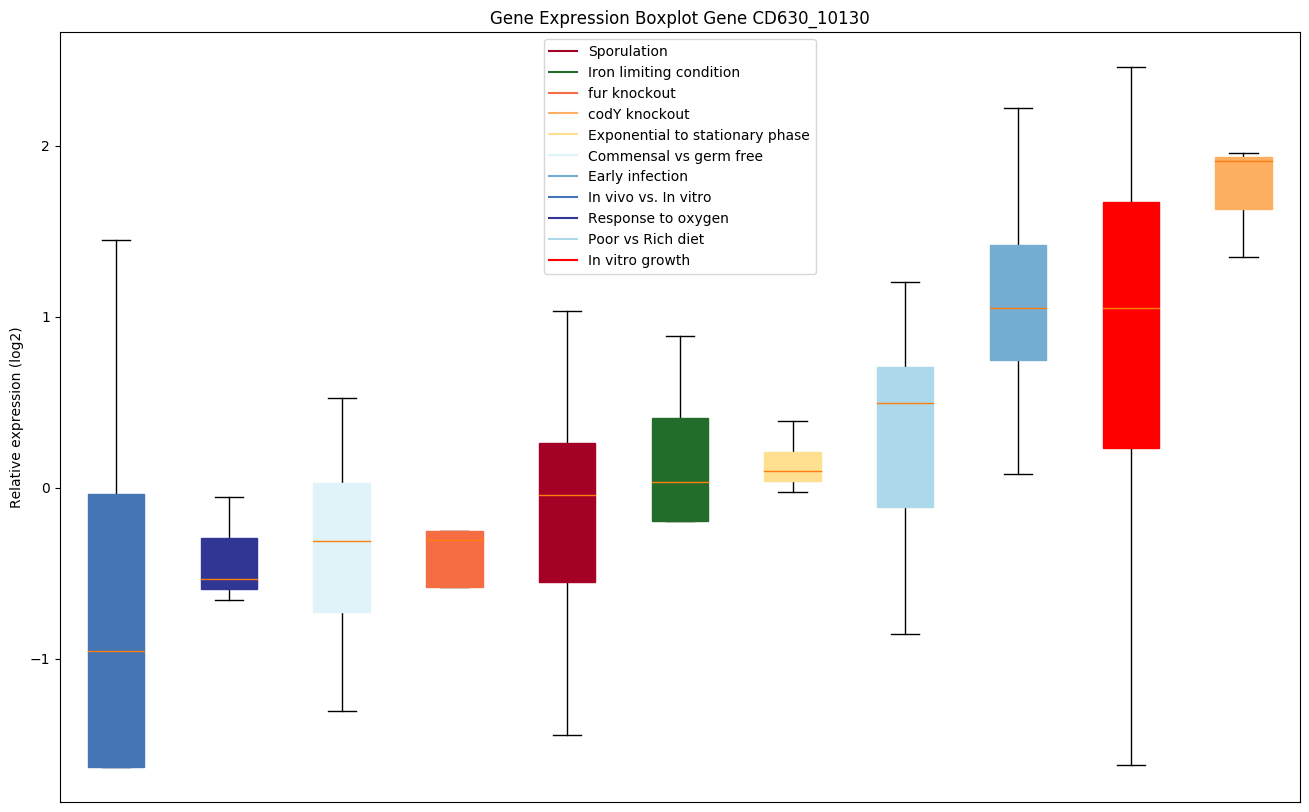 |
|||||
| CD630_24780 | Putative DNA uptake transporter |
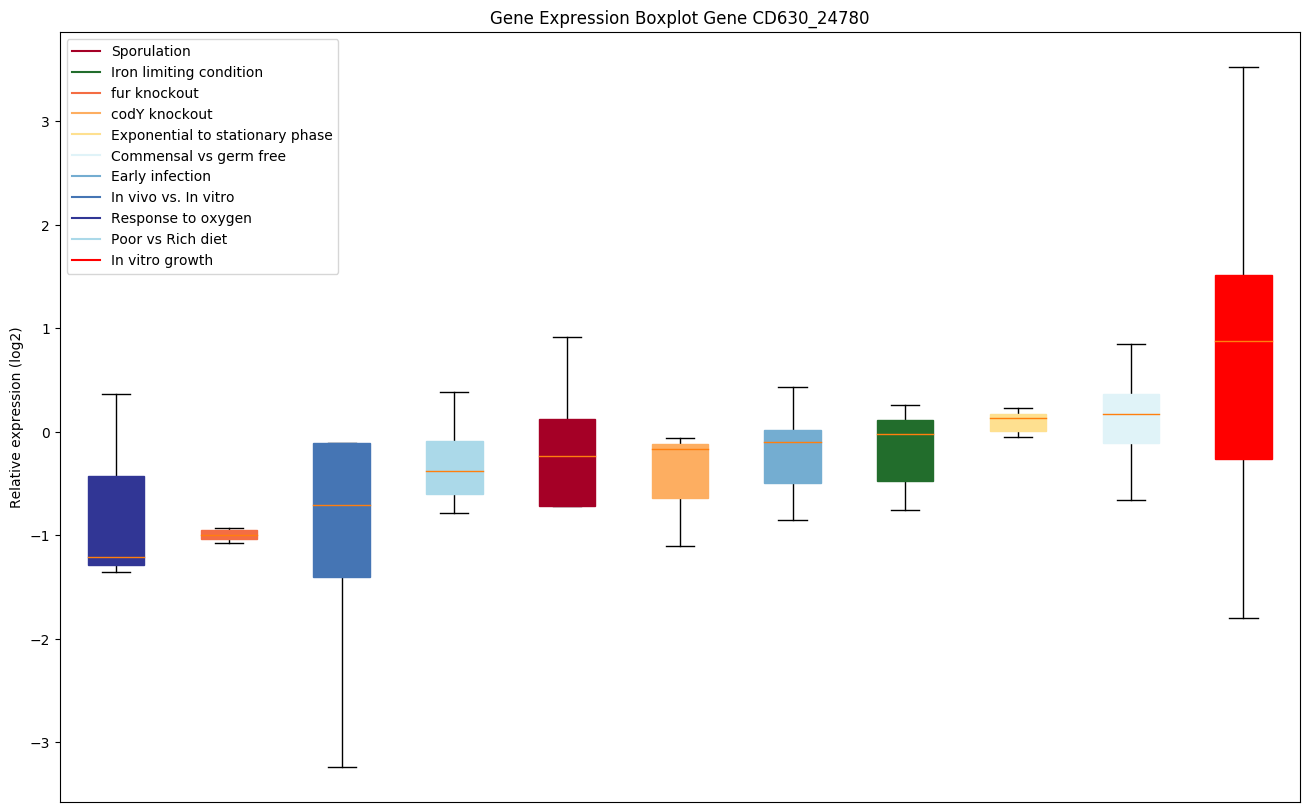 |
|||||
| CD630_25710 | Two-component response regulator |
 |
|||||
| CD630_34120 | uvrB | Excinuclease ABC subunit B | The UvrABC repair system catalyzes the recognition and processing of DNA lesions. A damage recognition complex composed of 2 UvrA and 2 UvrB subunits scans DNA for abnormalities. Upon binding of the UvrA(2)B(2) complex to a putative damaged site, the DNA wraps around one UvrB monomer. DNA wrap is dependent on ATP binding by UvrB and probably causes local melting of the DNA helix, facilitating insertion of UvrB beta-hairpin between the DNA strands. Then UvrB probes one DNA strand for the presence of a lesion. If a lesion is found the UvrA subunits dissociate and the UvrB-DNA preincision complex is formed. This complex is subsequently bound by UvrC and the second UvrB is released. If no lesion is found, the DNA wraps around the other UvrB subunit that will check the other stand for damage. |
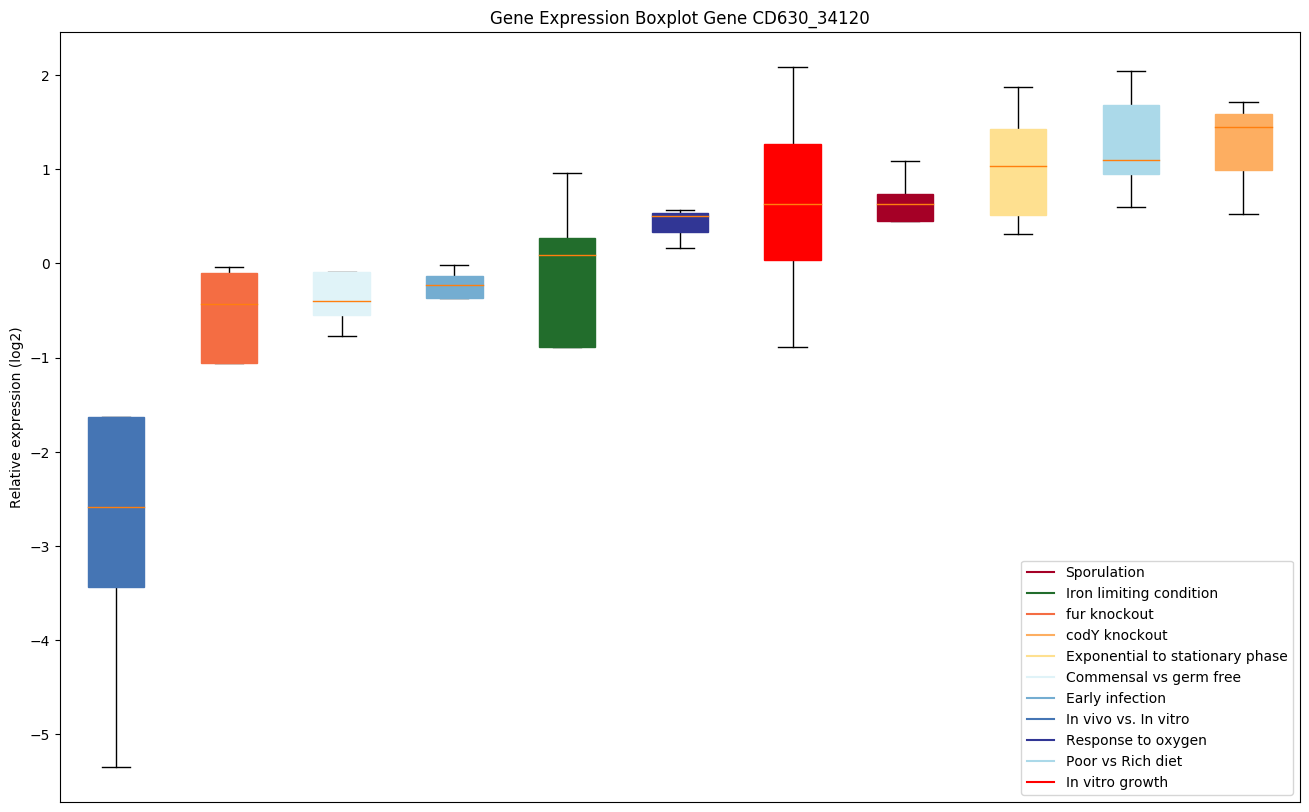 |
|||
| CD630_34100 | uvrC | Excinuclease ABC subunit C | The UvrABC repair system catalyzes the recognition and processing of DNA lesions. UvrC both incises the 5' and 3' sides of the lesion. The N-terminal half is responsible for the 3' incision and the C-terminal half is responsible for the 5' incision. |
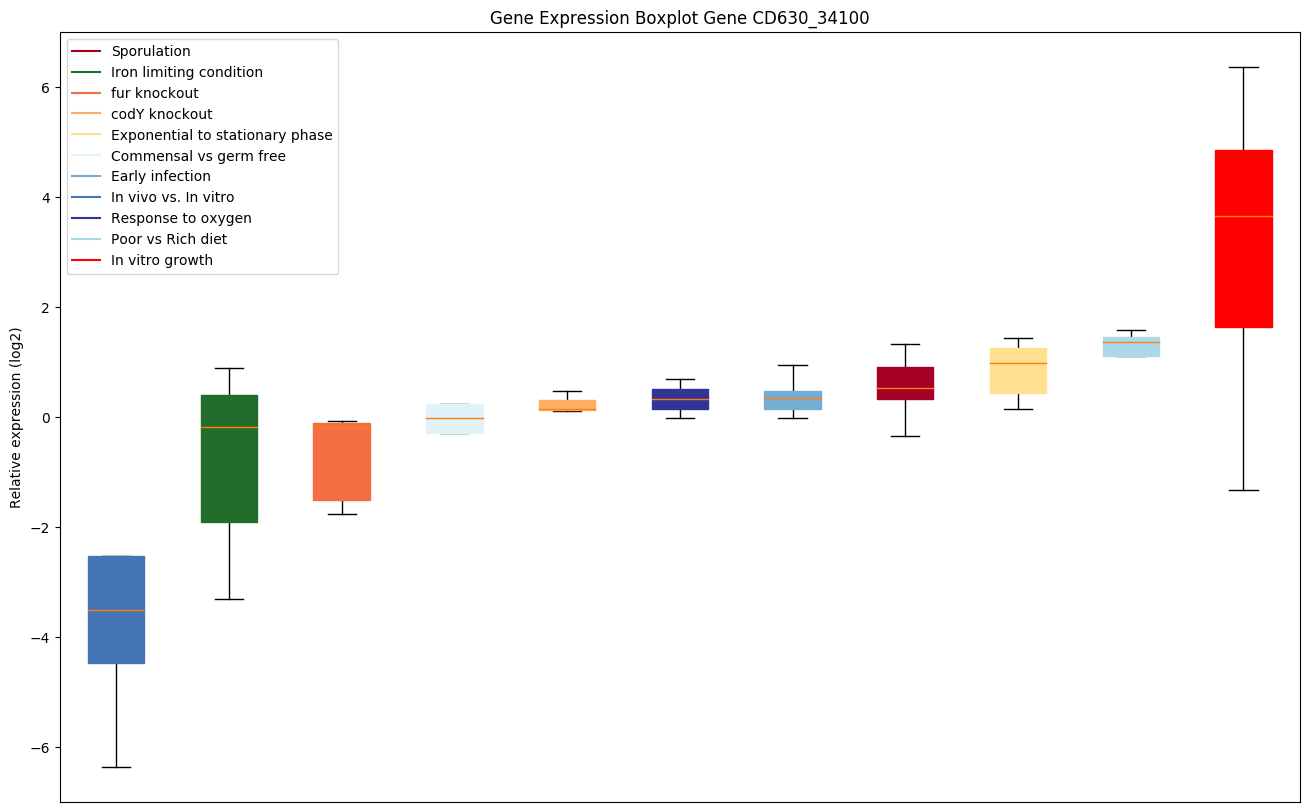 |
|||
| CD630_30450 | Putative ATPase, BadF/BadG/BcrA/BcrD type |
 |
|||||
| CD630_34090 | hprK | PTS system, HPr kinase/phosphorylase | Catalyzes the ATP- as well as the pyrophosphate-dependent phosphorylation of a specific serine residue in HPr, a phosphocarrier protein of the phosphoenolpyruvate-dependent sugar phosphotransferase system (PTS). HprK/P also catalyzes the pyrophosphate-producing, inorganic phosphate-dependent dephosphorylation (phosphorolysis) of seryl-phosphorylated HPr (P-Ser-HPr). The two antagonistic activities of HprK/P are regulated by several intracellular metabolites, which change their concentration in response to the absence or presence of rapidly metabolisable carbon sources (glucose, fructose, etc.) in the growth medium. Therefore, by controlling the phosphorylation state of HPr, HPrK/P is a sensor enzyme that plays a major role in the regulation of carbon metabolism and sugar transport: it mediates carbon catabolite repression (CCR), and regulates PTS-catalyzed carbohydrate uptake and inducer exclusion. |
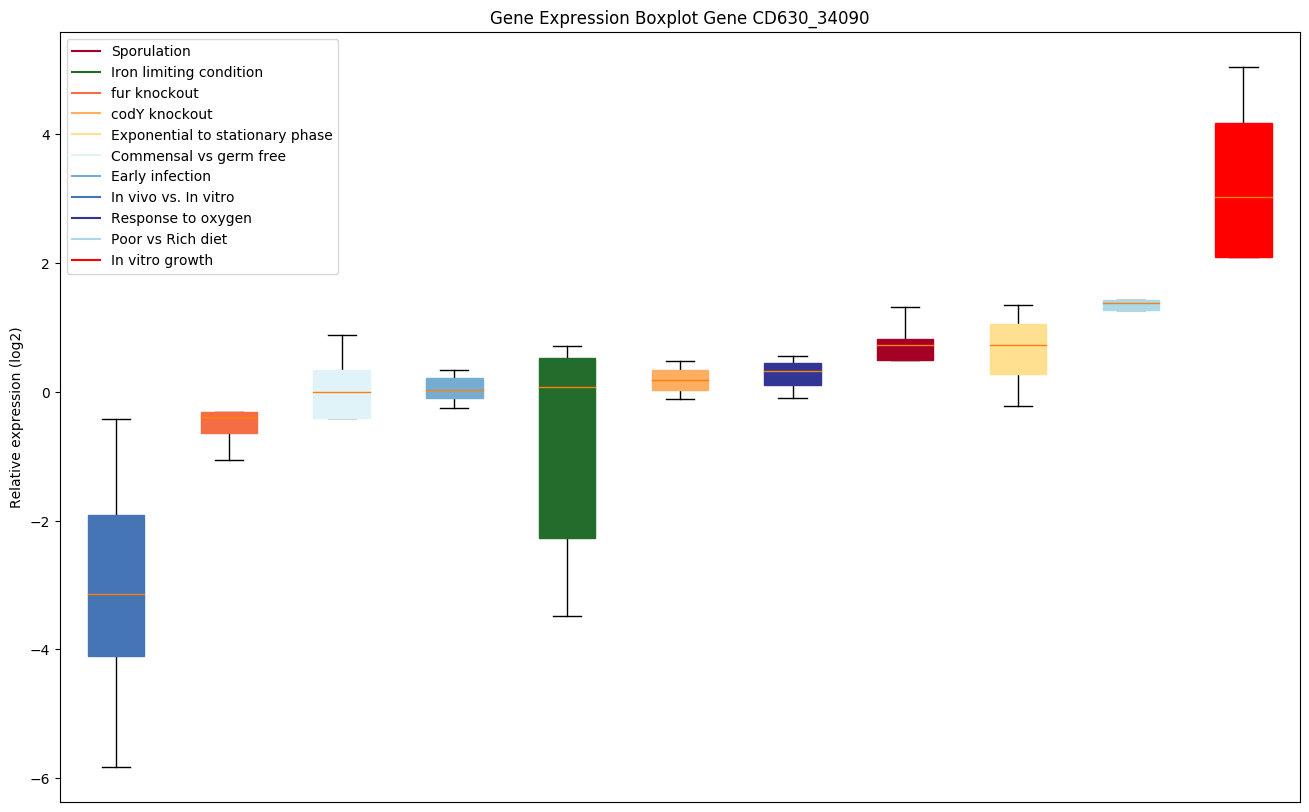 |
|||
| CD630_32170 | ABC-type transport system, ATP-binding protein |
 |
|||||
| CD630_22860 | Putative transporter |
 |
|||||
| CD630_12620 | rnhB | Ribonuclease HII (RNase HII) | Endonuclease that specifically degrades the RNA of RNA-DNA hybrids. |
 |
|||
| CD630_24740 | Putative DNA polymerase III, delta subunit | Yes |
 |
||||
| CD630_25240 | nadD | Nicotinate-nucleotide adenylyltransferase | Catalyzes the reversible adenylation of nicotinate mononucleotide (NaMN) to nicotinic acid adenine dinucleotide (NaAD). | Yes |
 |
||
| CD630_01910 | Putative DNA glycosylase |
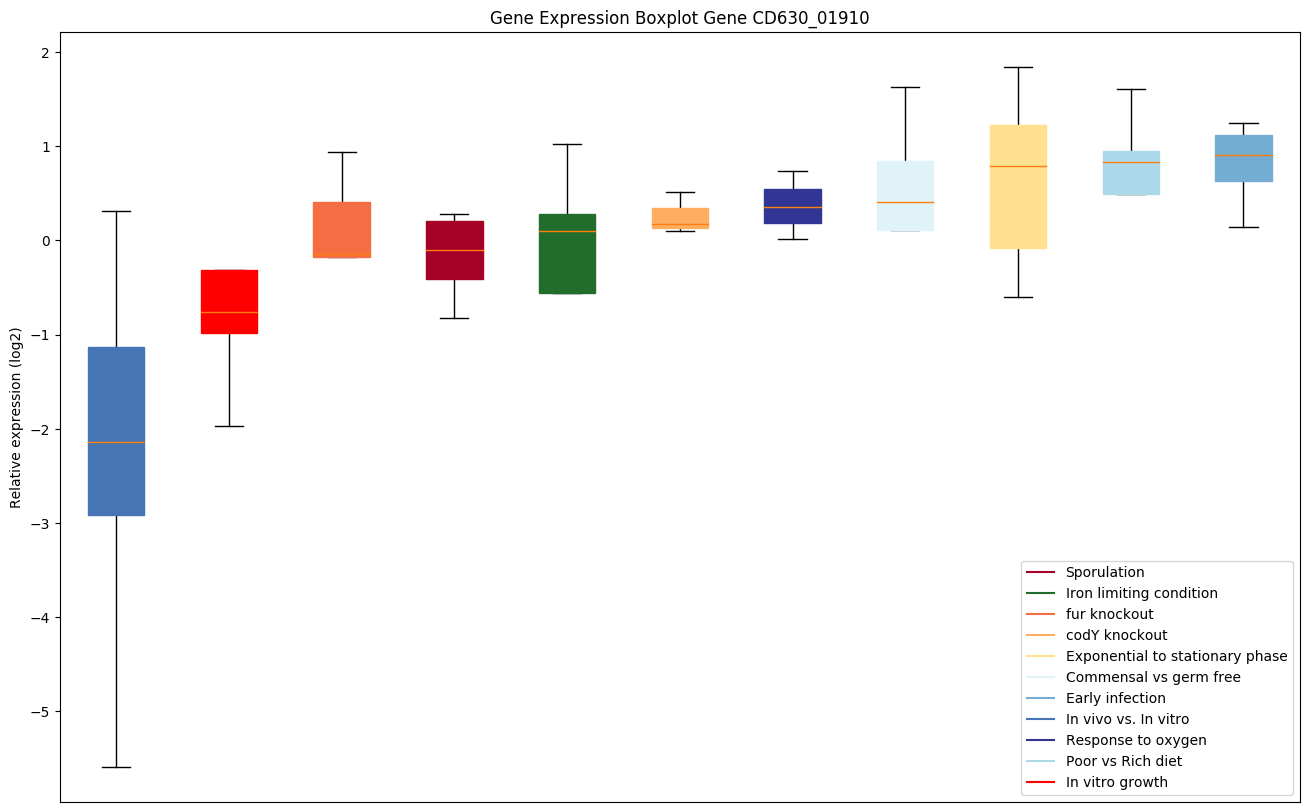 |
|||||
| CD630_24760 | Conserved hypothetical protein |
 |
|||||
| CD630_10140 | Two-component sensor histidine kinase |
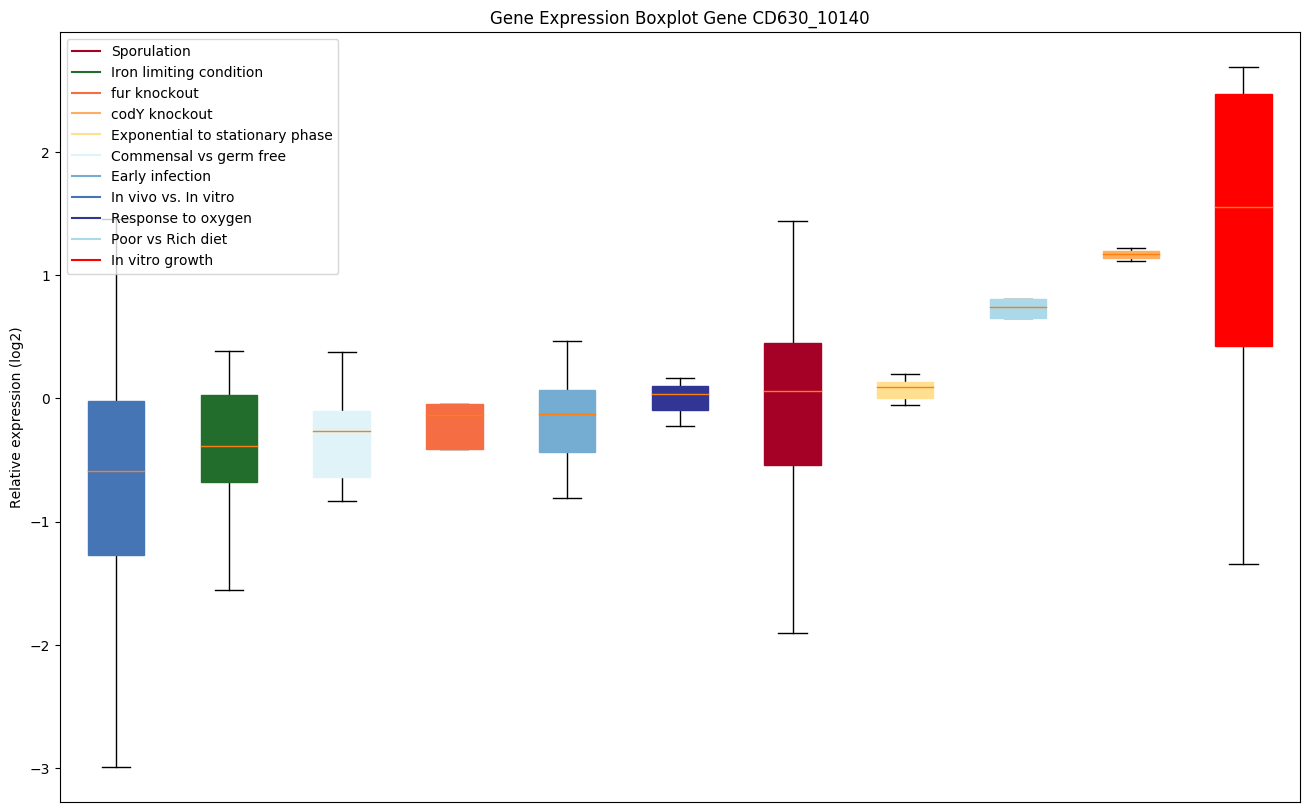 |
|||||
| CD630_12640 | Conserved hypothetical protein |
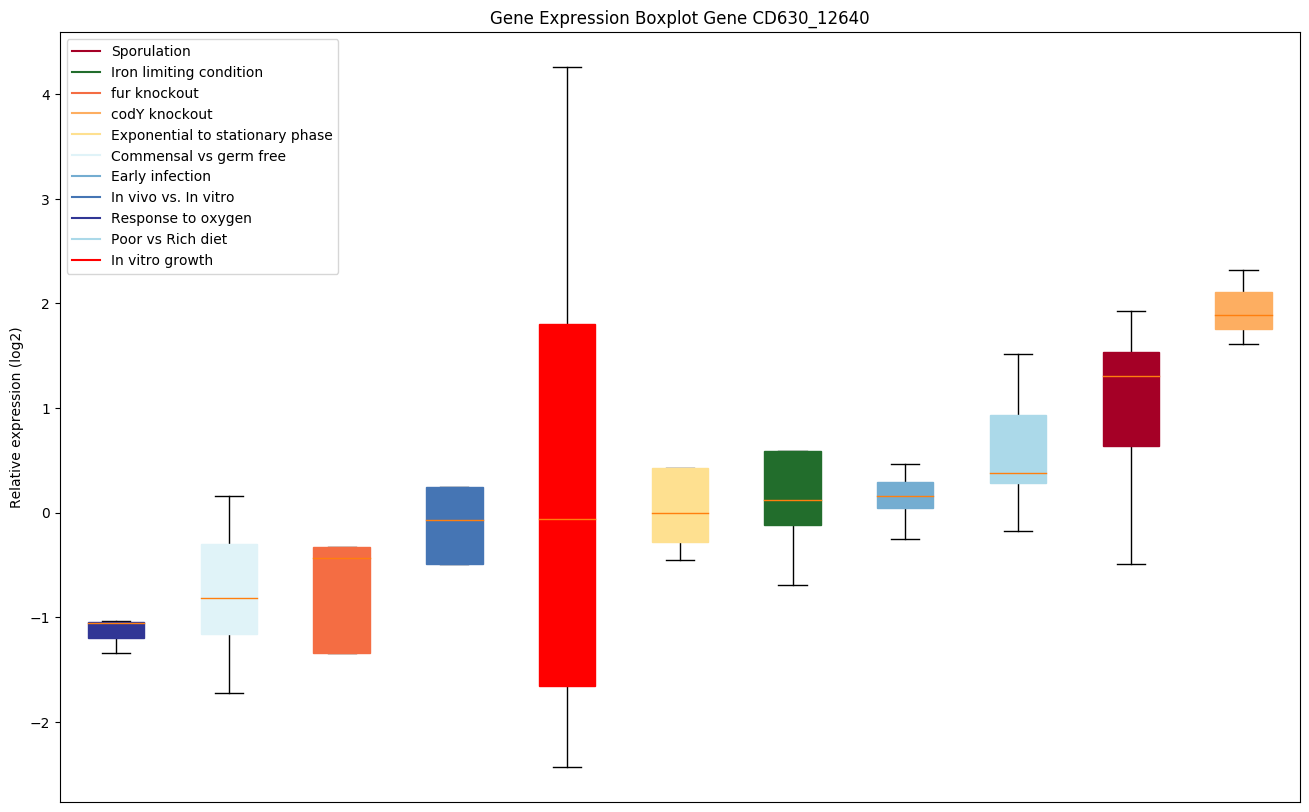 |
|||||
| CD630_25700 | Two-component sensor histidine kinase |
 |
|||||
| CD630_34110 | uvrA | Excinuclease ABC subunit A | The UvrABC repair system catalyzes the recognition and processing of DNA lesions. UvrA is an ATPase and a DNA-binding protein. A damage recognition complex composed of 2 UvrA and 2 UvrB subunits scans DNA for abnormalities. When the presence of a lesion has been verified by UvrB, the UvrA molecules dissociate. |
 |
|||
| CD630_25230 | Putative hydrolase |
 |