Module 147
Summary
| Residual | Gene Count | Condition Count | 1st Q Condition Block * | 5th Q Condition Block * |
|---|---|---|---|---|
| 0.34 | 21 | 65 | In vivo vs. In vitro | NA |
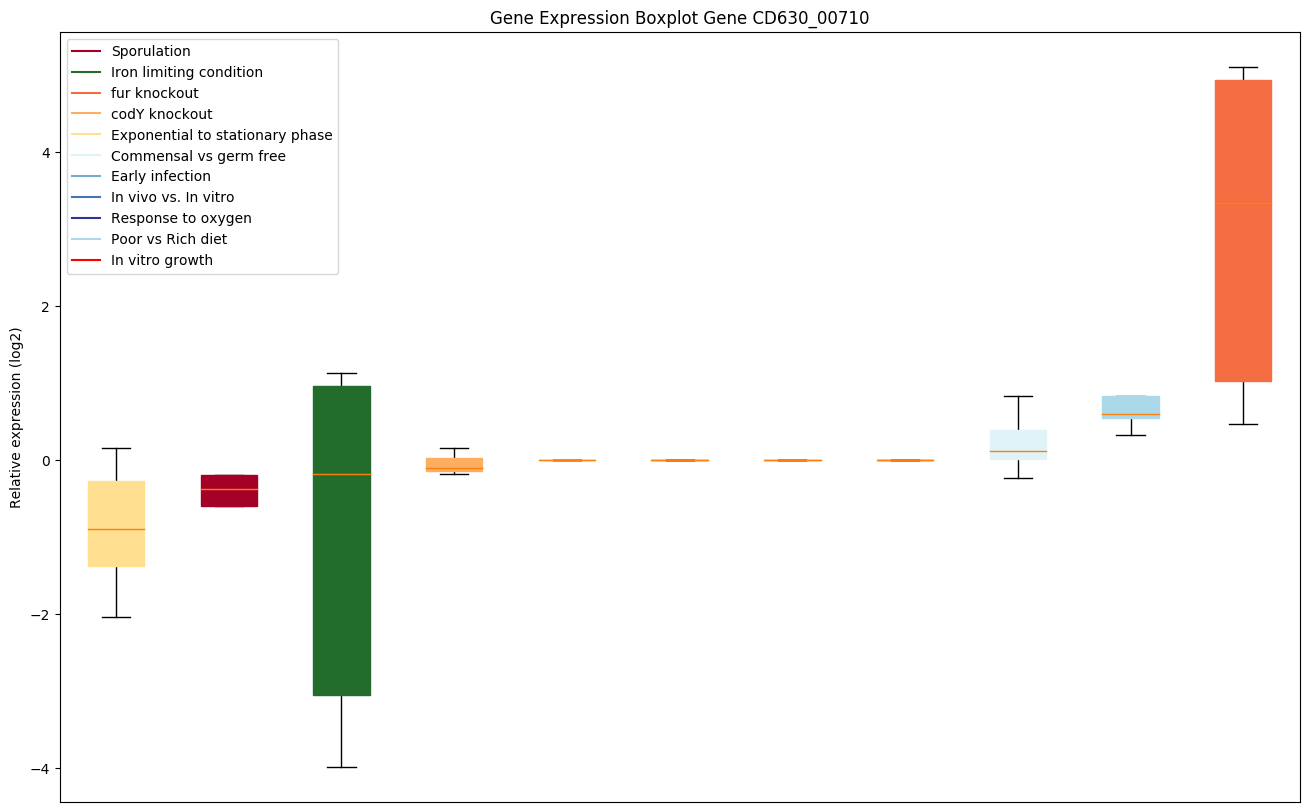
Bicluster expression profile
Expression of genes in subset of conditions included in bicluster on left side of red dashed line and out of bicluster on right of red dashed line. Each condition is represented as a boxplot, ordered by their median expression for the bicluster genes (smallest to largest) and colored according to condition blocks.
|
|
de novo identified motifs are listed below
| Motif 1 | Motif 1 e-value | Motif 2 | Motif 2 e-value | ||
|---|---|---|---|---|---|
| NA | NA | NA | NA |
Candidate transcriptional regulators for each module were determined with a linear regression based approach (Inferelator) and by evaluating the statistical significance of the overlap (Hypergeometric) between modules and putative TF and alternative sigma factor regulons (CcpA, CodY, Fur, PrdR, SigB, SigD, SigE, SigF, SigG, SigH, SigK, Spo0A) compiled from available literature.
For filtering high confidence influences, we used hypergeometric test adjusted p-value <=0.05 and minimum four genes in the overlap between the module and the TF regulon. The same thresholds were used for evaluating the functional enrichment.
1. Betas: correspond to the average coefficients of the Bayesian regressions between module expression and TF expression profiles. The values indicate the magnitude and direction (activation or repression for positive or negative values, respectively) of each TF-module interaction.
2. Confidence scores: indicate the likelihood of the TF-module interactions.
| TF | Module | Confidence Score | Beta |
|---|---|---|---|
|
Putative dinitrogenase iron-molybdenum cofactor |
147 | 0.95 | 0.8644 |
Candidate transcriptional regulators for each module were determined with a linear regression based approach (Inferelator) and by evaluating the statistical significance of the overlap (Hypergeometric) between modules and putative TF and alternative sigma factor regulons (CcpA, CodY, Fur, PrdR, SigB, SigD, SigE, SigF, SigG, SigH, SigK, Spo0A) compiled from available literature.
For filtering high confidence influences, we used hypergeometric test adjusted p-value <=0.05 and minimum four genes in the overlap between the module and the TF regulon. The same thresholds were used for evaluating the functional enrichment.
| TF | Module | Overlap | pvalue | Adjusted pvalue |
|---|---|---|---|---|
|
glnF RNA polymerase sigma-54 factor |
147 | 6 | 0.00 | 0.00 |
The module is significantly enriched with genes associated to the indicated functional terms
The enrichment was determined with hypergeometric test using the genome functional annotation compiled by Girinathan et al (2020).
| Module | Pathway | Overlap | pvalue | Adjusted pvalue |
|---|---|---|---|---|
| 147 | Stickland Fermentations | 5 | 0.000000 | 0.000011 |
| 147 | Fatty Acid Metabolism | 8 | 0.000000 | 0.000000 |
Genes that are included in this module
* "Gene essentiality is based on TnSeq Data from: Dembek M, Barquist L, Boinett CJ, et al. High-throughput analysis of gene essentiality and sporulation in Clostridium difficile. mBio. 2015;6(2):e02383. Published 2015 Feb 24.doi:10.1128/mBio.02383-14".
| Title | Short Name | Product | Function | Essentiality * | in vivo Essentiality | Rich broth Essentiality | Expression |
|---|---|---|---|---|---|---|---|
| CD630_03960 | hadI | Activator of 2-hydroxyisocaproyl-CoA dehydratase |
 |
||||
| CD630_04010 | etfA1 | Alpha-subunit of electron transfer flavoprotein |
 |
||||
| CD630_11780 | plsX | Fatty acid/phospholipid synthesis protein PlsX | Catalyzes the reversible formation of acyl-phosphate (acyl-PO(4)) from acyl-[acyl-carrier-protein] (acyl-ACP). This enzyme utilizes acyl-ACP as fatty acyl donor, but not acyl-CoA. | Yes | Yes | Yes |
 |
| CD630_11800 | fabK | Enoyl-(Acyl-carrier-protein) reductase II | Yes | Yes | Yes |
 |
|
| CD630_03950 | hadA | Isocaprenoyl-CoA:2-hydroxyisocaproateCoA-transferase |
 |
||||
| CD630_11770 | fapR | Transcriptional regulator, DeoR family (Fattyacid and phospholipid biosynthesis regulator) | Yes |
 |
|||
| CD630_11810 | fabD | Malonyl CoA-acyl carrier protein transacylase(MCT) | Yes | Yes | Yes |
 |
|
| CD630_11790 | fabH | 3-oxoacyl-[acyl-carrier-protein] synthase 3 | Catalyzes the condensation reaction of fatty acid synthesis by the addition to an acyl acceptor of two carbons from malonyl-ACP. Catalyzes the first condensation reaction which initiates fatty acid synthesis and may therefore play a role in governing the total rate of fatty acid production. Possesses both acetoacetyl-ACP synthase and acetyl transacylase activities. Its substrate specificity determines the biosynthesis of branched-chain and/or straight-chain of fatty acids. | Yes | Yes | Yes |
 |
| CD630_03990 | acdB | Acyl-CoA dehydrogenase, short-chain specific |
 |
||||
| CD630_00941 | rpmJ | 50S ribosomal protein L36 | Yes |
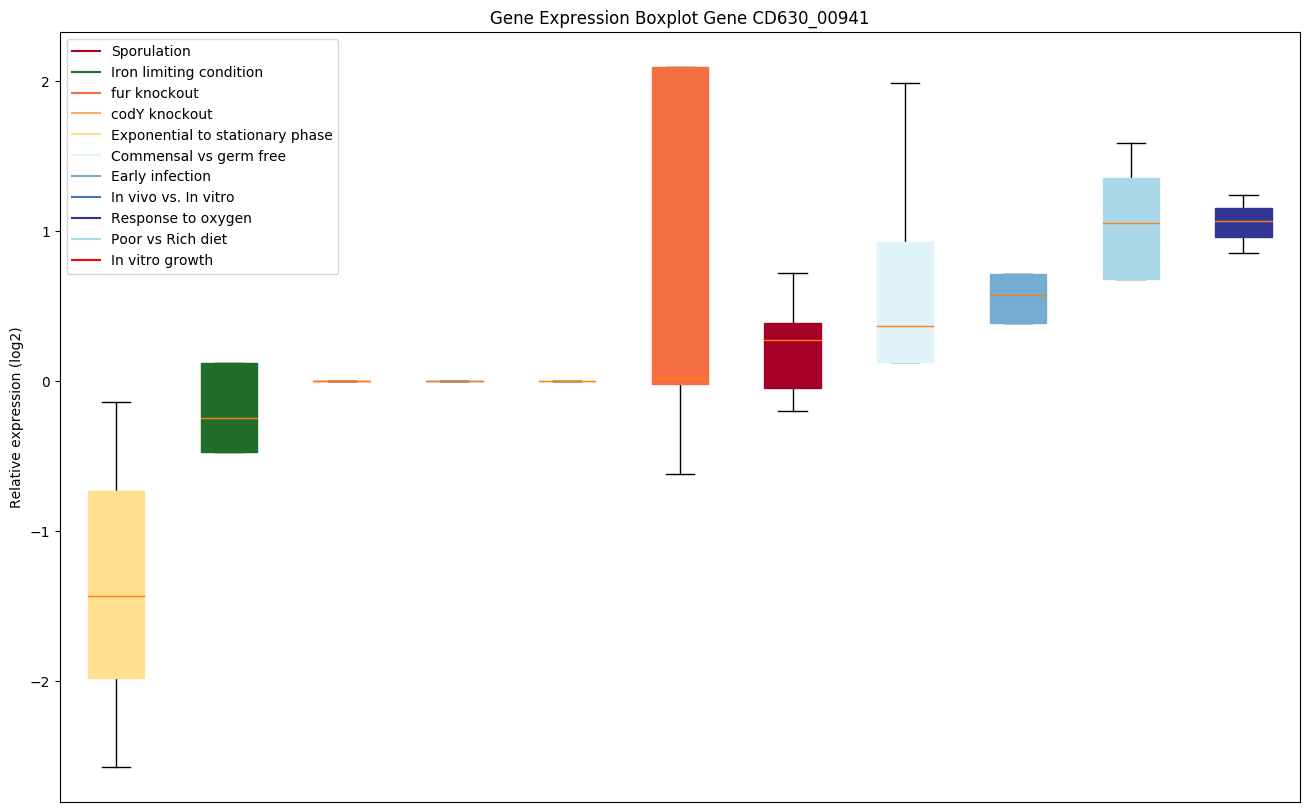 |
|||
| CD630_09231 | Putative phage protein |
 |
|||||
| CD630_00881 | rpmD | 50S ribosomal protein L30 | Yes |
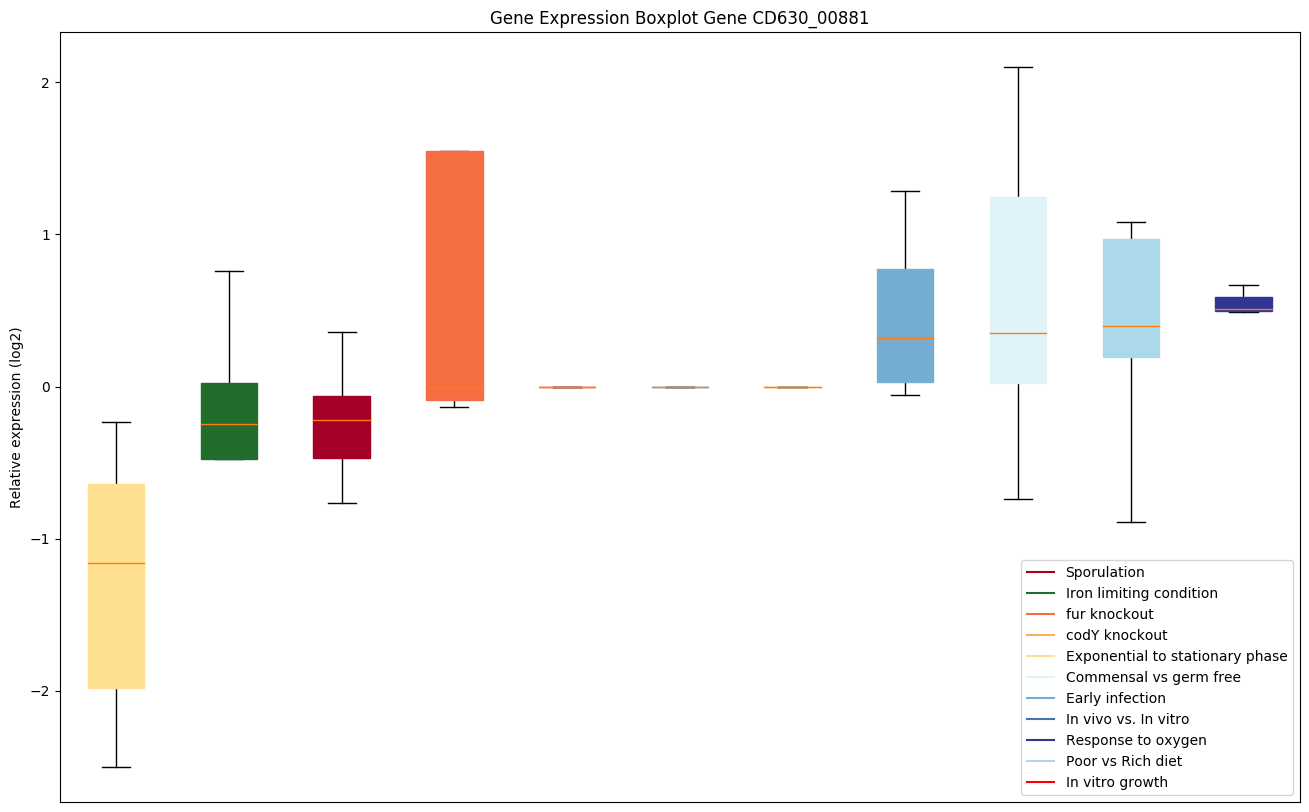 |
|||
| CD630_03980 | hadC | Subunit of oxygen-sensitive2-hydroxyisocaproyl-CoA dehydratase C |
 |
||||
| CD630_11840 | fabF | 3-oxoacyl-[acyl-carrier-protein] synthase 2 | Catalyzes the condensation reaction of fatty acid synthesis by the addition to an acyl acceptor of two carbons from malonyl-ACP. | Yes | Yes | Yes |
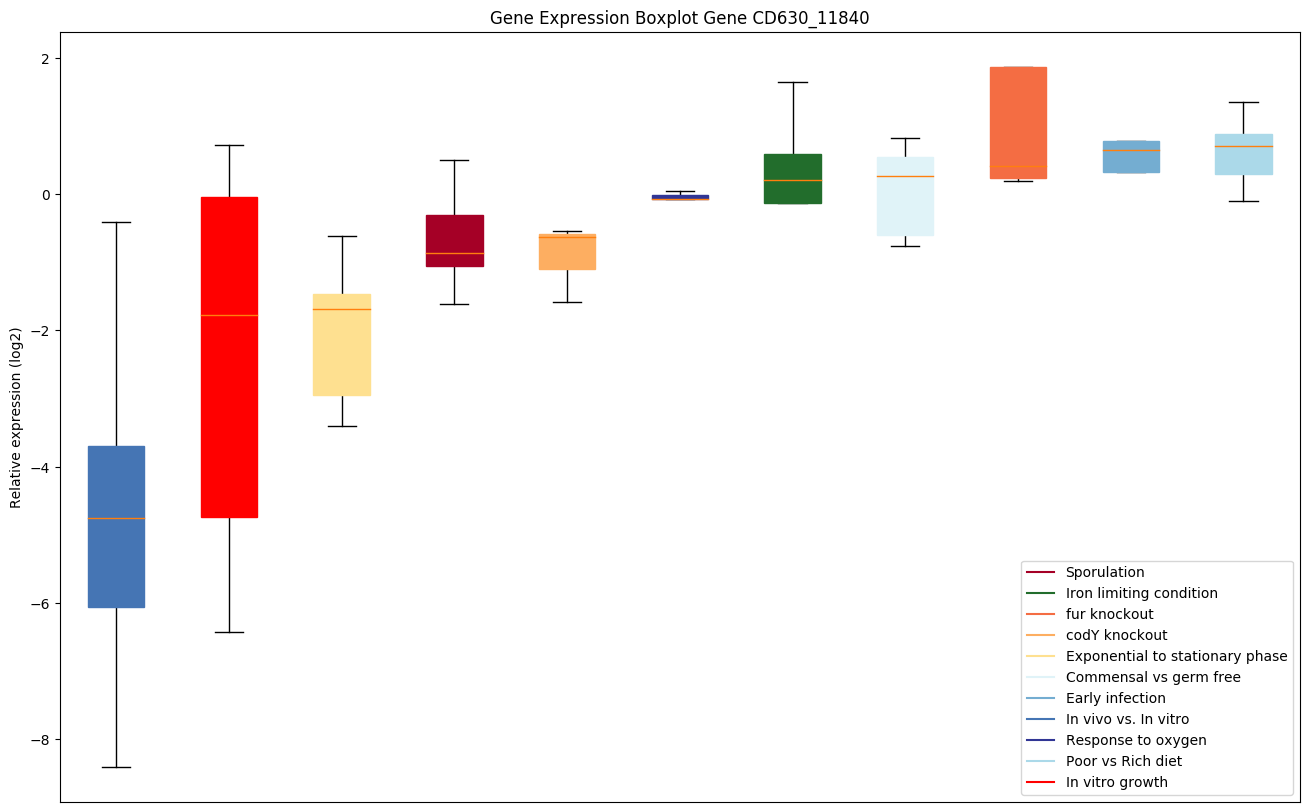 |
| CD630_00841 | rpsZ | 30S ribosomal protein S14 type Z | Binds 16S rRNA, required for the assembly of 30S particles and may also be responsible for determining the conformation of the 16S rRNA at the A site. | Yes |
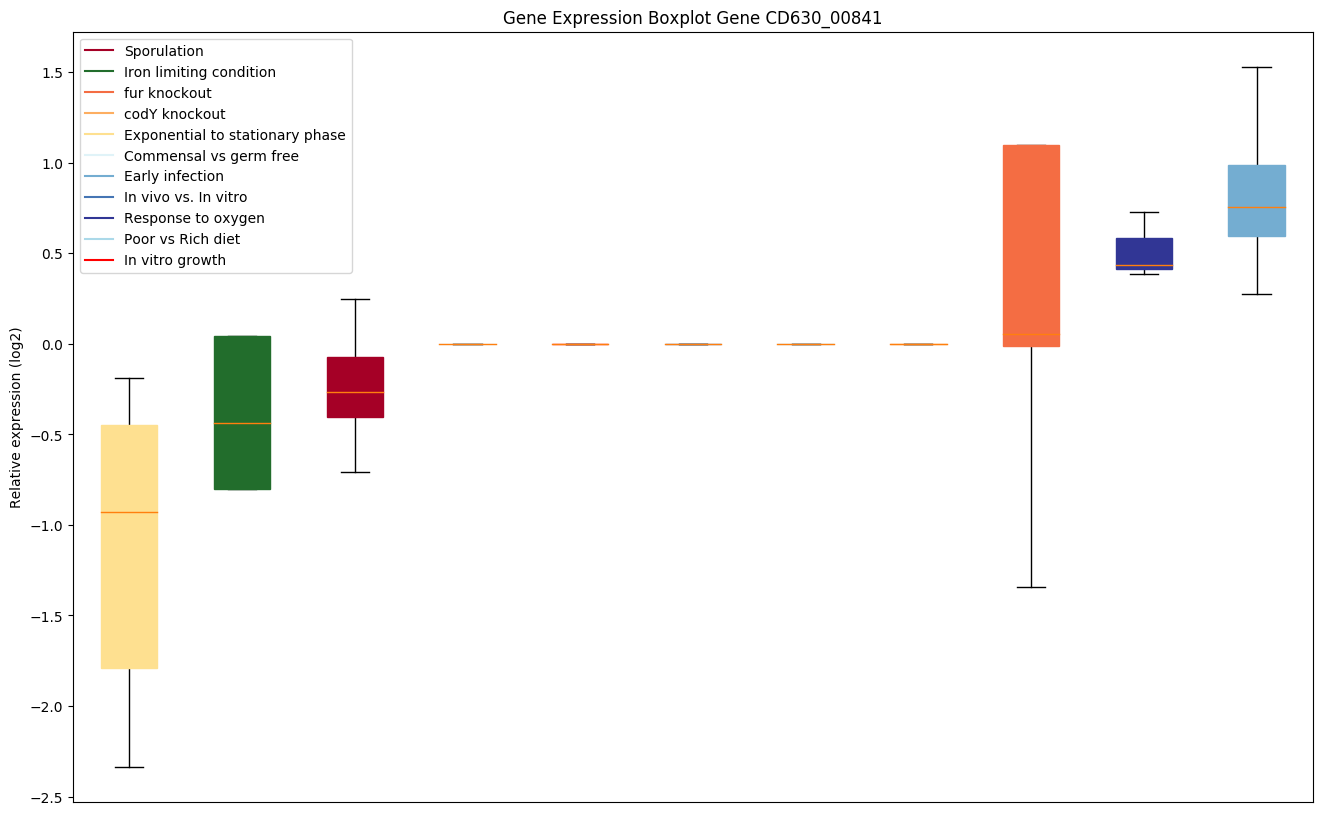 |
||
| CD630_11820 | fabG | 3-oxoacyl-[acyl-carrier protein] reductase(3-ketoacyl-acyl carrier protein reductase) | Catalyzes the NADPH-dependent reduction of beta-ketoacyl-ACP substrates to beta-hydroxyacyl-ACP products, the first reductive step in the elongation cycle of fatty acid biosynthesis. | Yes |
 |
||
| CD630_11830 | acpP | Acyl carrier protein (ACP) | Carrier of the growing fatty acid chain in fatty acid biosynthesis. | Yes |
 |
||
| CD630_03970 | hadB | Subunit of oxygen-sensitive2-hydroxyisocaproyl-CoA dehydratase B |
 |
||||
| CD630_04000 | etfB1 | Beta-subunit of electron transfer flavoprotein |
 |
||||
| CD630_00710 | tuf | Elongation factor EFTu/EF1A (EF-Tu) | This protein promotes the GTP-dependent binding of aminoacyl-tRNA to the A-site of ribosomes during protein biosynthesis. | Yes |
 |
||
| CD630_00801 | rpmC | 50S ribosomal protein L29 | Yes |
 |
