Module 55 Residual: 0.58
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0055 | v02 | 0.58 | -13.54 |
| Rv0566c UPF0234 protein YajQ | Rv0567 O-methyltransferase |
| Rv0759c HIT family protein | Rv0831c |
| Rv1018c N-acetylglucosamine-1-phosphate uridyltransferase (EC 2.7.7.23) / Glucosamine-1-phosphate N-acetyltransferase (EC 2.3.1.157) | Rv1019 Transcriptional regulator, TetR family |
| Rv1020 Transcription-repair coupling factor | Rv1021 Nucleoside triphosphate pyrophosphohydrolase MazG (EC 3.6.1.8) |
| Rv1292 Arginyl-tRNA synthetase (EC 6.1.1.19) | Rv1293 Diaminopimelate decarboxylase (EC 4.1.1.20) |
| Rv1295 Threonine synthase (EC 4.2.3.1) | Rv1746 Probable serine/threonine-protein kinase pknF (EC 2.7.11.1) |
| Rv2272 Possible membrane protein | Rv2273 Possible membrane protein |
| Rv2289 CDP-diacylglycerol pyrophosphatase (EC 3.6.1.26) | Rv2290 Probable conserved lipoprotein lppOb |
|
GO:0005829 | cytosol |
|
GO:0005886 | plasma membrane |
|
GO:0005618 | cell wall |
|
GO:0008150 | biological_process |
|
GO:0000287 | magnesium ion binding |
|
GO:0003977 | UDP-N-acetylglucosamine diphosphorylase activity |
|
GO:0019134 | glucosamine-1-phosphate N-acetyltransferase activity |
|
GO:0040007 | growth |
|
GO:0070569 | uridylyltransferase activity |
|
GO:0003700 | sequence-specific DNA binding transcription factor activity |
|
GO:0008026 | ATP-dependent helicase activity |
|
GO:0003676 | nucleic acid binding |
|
GO:0004386 | helicase activity |
|
GO:0005524 | ATP binding |
|
GO:0004814 | arginine-tRNA ligase activity |
|
GO:0008836 | diaminopimelate decarboxylase activity |
|
GO:0042803 | protein homodimerization activity |
|
GO:0051289 | protein homotetramerization |
|
GO:0004795 | threonine synthase activity |
|
GO:0009088 | threonine biosynthetic process |
|
GO:0030170 | pyridoxal phosphate binding |
|
GO:0000917 | barrier septum assembly |
|
GO:0001558 | regulation of cell growth |
|
GO:0004672 | protein kinase activity |
|
GO:0004674 | protein serine/threonine kinase activity |
|
GO:0005515 | protein binding |
|
GO:0005887 | integral to plasma membrane |
|
GO:0010827 | regulation of glucose transport |
|
GO:0043085 | positive regulation of catalytic activity |
|
GO:0043086 | negative regulation of catalytic activity |
|
GO:0051055 | negative regulation of lipid biosynthetic process |
|
GO:0008715 | CDP-diacylglycerol diphosphatase activity |
|
GO:0005576 | extracellular region |



















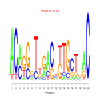
Discussion