Module 35 Residual: 0.52
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0035 | v02 | 0.52 | -13.26 |
| Rv0389 Phosphoribosylglycinamide formyltransferase 2 (EC 2.1.2.-) | Rv0751c 3-hydroxyisobutyrate dehydrogenase (EC 1.1.1.31) |
| Rv0752c Acyl-CoA dehydrogenase, short-chain specific (EC 1.3.99.2) | Rv0753c Methylmalonate-semialdehyde dehydrogenase (EC 1.2.1.27) |
| Rv0754 PE-PGRS FAMILY PROTEIN | Rv0775 Transcriptional regulator, TetR family |
| Rv1997 Probable cation-transporting ATPase F (EC 3.6.3.-) | Rv2003c |
| Rv2004c | Rv2159c |
| Rv2629 | Rv2630 Archease |
| Rv2665 | Rv2964 Formyltetrahydrofolate deformylase (EC 3.5.1.10) |
| Rv3916c Conserved hypothetical protein |
| Module GO Term Enrichment | Descriptions |
|---|---|
| GO:0016742 | hydroxymethyl-, formyl- and related tran... |
|
GO:0005829 | cytosol |
|
GO:0008442 | 3-hydroxyisobutyrate dehydrogenase activity |
|
GO:0004085 | butyryl-CoA dehydrogenase activity |
|
GO:0004491 | methylmalonate-semialdehyde dehydrogenase (acylating) activity |
|
GO:0005886 | plasma membrane |
|
GO:0005618 | cell wall |
|
GO:0005887 | integral to plasma membrane |
|
GO:0071500 | cellular response to nitrosative stress |
|
GO:0046812 | host cell surface binding |
|
GO:0001666 | response to hypoxia |
|
GO:0008864 | formyltetrahydrofolate deformylase activity |
|
GO:0051409 | response to nitrosative stress |

















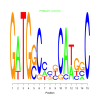
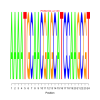
Discussion