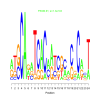
Module 82 Residual: 0.54
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0082 | v02 | 0.54 | -8.78 |
| Rv0950c Phage peptidoglycan binding endopeptidase | Rv1172c PE-PGRS FAMILY PROTEIN |
| Rv1617 Pyruvate kinase (EC 2.7.1.40) | Rv1618 Acyl-CoA thioesterase II (EC 3.1.2.-) |
| Rv2357c Glycyl-tRNA synthetase (EC 6.1.1.14) | Rv2430c PPE family protein |
| Rv2443 | Rv3377c Ent-copalyl diphosphate synthase |
| Rv3378c | Rv3646c DNA topoisomerase I (EC 5.99.1.2) |
| Rv3722c Aspartate transaminase (EC 2.6.1.1) | Rv2264c Possible molecular chaperone DnaK |
| Rv2289 CDP-diacylglycerol pyrophosphatase (EC 3.6.1.26) |
|
GO:0004743 | pyruvate kinase activity |
|
GO:0005829 | cytosol |
|
GO:0005886 | plasma membrane |
|
GO:0040007 | growth |
|
GO:0004820 | glycine-tRNA ligase activity |
|
GO:0005515 | protein binding |
|
GO:0005576 | extracellular region |
|
GO:0035439 | halimadienyl-diphosphate synthase activity |
|
GO:0035440 | tuberculosinol biosynthetic process |
|
GO:0052572 | response to host immune response |
|
GO:0016102 | diterpenoid biosynthetic process |
|
GO:0016791 | phosphatase activity |
|
GO:0003917 | DNA topoisomerase type I activity |
|
GO:0000287 | magnesium ion binding |
|
GO:0005618 | cell wall |
|
GO:0008715 | CDP-diacylglycerol diphosphatase activity |
















Discussion