Module 439 Residual: 0.56
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0439 | v02 | 0.56 | -18.79 |
| Rv0432 Superoxide dismutase [Cu-Zn] precursor (EC 1.15.1.1) | Rv3031 Glycogen branching enzyme, GH-57-type, archaeal (EC 2.4.1.18) |
| Rv0433 Carboxylate-amine ligase | Rv1098c Fumarate hydratase class II (EC 4.2.1.2) |
| Rv2000 | Rv2001 Acyl-ACP thioesterase, FatA |
| Rv2002 3-alpha-(or 20-beta)-hydroxysteroid dehydrogenase (EC 1.1.1.53) | Rv2271 |
| Rv2299c Chaperone protein HtpG | Rv2357c Glycyl-tRNA synthetase (EC 6.1.1.14) |
| Rv3029c Electron transfer flavoprotein, beta subunit | Rv3030 Methyltransferase (EC 2.1.1.-) |
| Rv3715c Recombination protein RecR | Rv3716c |
| Rv3722c Aspartate transaminase (EC 2.6.1.1) |
|
GO:0004784 | superoxide dismutase activity |
|
GO:0005886 | plasma membrane |
|
GO:0006979 | response to oxidative stress |
|
GO:0033194 | response to hydroperoxide |
|
GO:0045454 | cell redox homeostasis |
|
GO:0052059 | evasion or tolerance by symbiont of host-produced reactive oxygen species |
|
GO:0052060 | evasion or tolerance by symbiont of host-produced nitric oxide |
|
GO:0075136 | response to host |
|
GO:0046872 | metal ion binding |
|
GO:0003844 | 1,4-alpha-glucan branching enzyme activity |
|
GO:0005576 | extracellular region |
|
GO:0040007 | growth |
|
GO:0004333 | fumarate hydratase activity |
|
GO:0005829 | cytosol |
|
GO:0005618 | cell wall |
|
GO:0008202 | steroid metabolic process |
|
GO:0047044 | androstan-3-alpha,17-beta-diol dehydrogenase activity |
|
GO:0005524 | ATP binding |
|
GO:0006457 | protein folding |
|
GO:0051082 | unfolded protein binding |
|
GO:0071451 | cellular response to superoxide |
|
GO:0004820 | glycine-tRNA ligase activity |
|
GO:0009055 | electron carrier activity |
|
GO:0006118 | electron transport |
|
GO:0006281 | DNA repair |
|
GO:0003676 | nucleic acid binding |
|
GO:0006310 | DNA recombination |
|
GO:0006304 | DNA modification |


















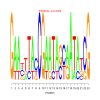
Discussion