Regulatory Modules
Displaying 51 - 60 of 170Desulfovibrio vulgaris Hildenborough
| Title | Motif 1 | Motif 2 | Residual | Bicluster Genes | Selected GO Terms |
|---|---|---|---|---|---|
| bicluster_0130 |
1900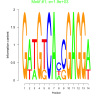 |
2500 |
0.38 | DVU0365, DVU1068, DVU1829, DVU2159, DVU2161, DVU2266, DVUA0009, DVU0375, DVU0497, DVU0887, DVU1490, DVU3129 | nitrogen cycle metabolic process, hydrolase activity, acting on glycosyl b... |
| bicluster_0213 |
6100 |
24000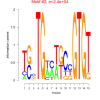 |
0.39 | DVU0183, DVU1103, DVU2731, DVU0039, DVU1713, DVU2217, DVU2559, DVU2604, DVU2689, DVU2830, DVU3338, DVU3339, DVUA0030 | N-acetyltransferase activity, zinc ion binding, N-acyltransferase activity, transaminase activity, acetyltransferase activity |
| bicluster_0088 |
980 |
2600 |
0.39 | DVU0106, DVU0321, DVU0945, DVU1027, DVU1406, DVU1652, DVU1778, DVU2372, DVU2607, DVU2608, DVU3085, DVU3088, DVU2650, DVU3149 | IMP biosynthetic process, 'de novo' IMP biosynthetic process, IMP metabolic process, purine nucleotide biosynthetic process, purine nucleoside monophosphate biosynth... |
| bicluster_0098 |
210 |
19000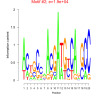 |
0.39 | DVU0280, DVU0281, DVU0669, DVU0817, DVU0954, DVU1688, DVU1690, DVU1793, DVU1796, DVU2373, DVU2569, DVUA0074, DVU1798, DVU1809, DVU1931 | cell envelope organization, Gram-negative-bacterium-type cell outer ..., membrane biogenesis, single-organism membrane organization, membrane organization |
| bicluster_0259 |
5700 |
2000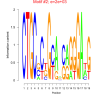 |
0.39 | DVU2127, DVU0888, DVU1107, DVU2120, DVU2129, DVU2151, DVU2157, DVU2191, DVU2194, DVU3306 | phosphorelay sensor kinase activity, protein kinase activity, protein histidine kinase activity, receptor activity, phosphotransferase activity, nitrogenous... |
| bicluster_0168 |
0.00079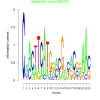 |
0.025 |
0.39 | DVU1105, DVU1695, DVU2596, DVU2598, DVU2600, DVU2623, DVU2705, DVU1542, DVU1543, DVU2601, DVU2602, DVU2632, DVU2664, DVU2701, DVU2704 | |
| bicluster_0171 |
120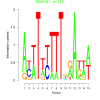 |
17000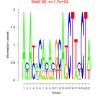 |
0.39 | DVU0698, DVU1075, DVU1301, DVU1427, DVU2200, DVU0817, DVU0886, DVU1778, DVU1849, DVU2179, DVU2215, DVU2220 | racemase and epimerase activity, racemase and epimerase activity, acting ..., endonuclease activity, cation transmembrane transporter activit... |
| bicluster_0036 |
28 |
1500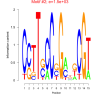 |
0.39 | DVU0081, DVU0083, DVU0316, DVU0484, DVU0494, DVU0514, DVU0636, DVU1433, DVU1940, DVU2663, DVU2865, DVU3034, DVU3106, DVU3387, DVU0486, DVU2622 | cofactor binding, ion binding, pyridoxal phosphate binding, FMN binding |
| bicluster_0336 |
950 |
660 |
0.39 | DVU0968, DVU0506, DVU0787, DVU0788, DVU0967, DVU1532, DVU1671, DVU1673, DVU1776, DVU1935, DVU2036, DVU2374, DVU2375, DVU3069, DVU3164, DVU3165 | cell morphogenesis, anatomical structure morphogenesis, developmental process, cellular component morphogenesis, single-organism developmental process |
| bicluster_0038 |
47 |
21000 |
0.39 | DVU0031, DVU2732, DVU2829, DVU2830, DVU2848, DVU2852, DVU2854, DVU2856, DVUA0014, DVUA0066, DVU0180, DVU0636, DVU2868 | enzyme regulator activity |

