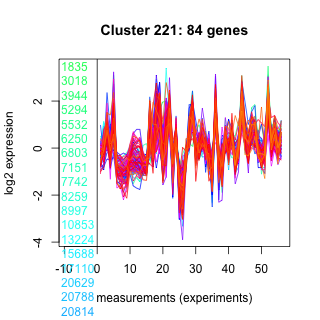
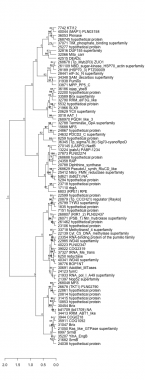
Thaps_hclust_0221 Hierarchical Clustering
Thalassiosira pseudonana
| GO ID | Go Term | p-value | q-value | Cluster |
|---|---|---|---|---|
| GO:0006177 | GMP biosynthesis | 0.0000846 | 0.306162 | Thaps_hclust_0221 |
| GO:0008033 | tRNA processing | 0.155103 | 1 | Thaps_hclust_0221 |
| GO:0016071 | mRNA metabolism | 0.00292321 | 1 | Thaps_hclust_0221 |
| GO:0009401 | phosphoenolpyruvate-dependent sugar phosphotransferase system | 0.155103 | 1 | Thaps_hclust_0221 |
| GO:0006260 | DNA replication | 0.0040006 | 1 | Thaps_hclust_0221 |
| GO:0006413 | translational initiation | 0.201373 | 1 | Thaps_hclust_0221 |
| GO:0006352 | transcription initiation | 0.00618548 | 1 | Thaps_hclust_0221 |
| GO:0006306 | DNA methylation | 0.238031 | 1 | Thaps_hclust_0221 |
| GO:0006106 | fumarate metabolism | 0.00930329 | 1 | Thaps_hclust_0221 |
| GO:0006810 | transport | 0.243952 | 1 | Thaps_hclust_0221 |
|
1835 : hypothetical protein |
21682 : SrmB |
27873 : PLN02274 |
36345 : (Tp_sigma70.3b) Sig70-cyanoRpoD |
|
3018 : AAT_I |
21933 : RNA_pol_I_A49 superfamily |
28189 : (HSP70_3) PTZ00009 |
37071 : TIM_phosphate_binding superfamily |
|
3944 : COG0218 |
21966 : SLX9 |
28441 : eIF-3c_N superfamily |
37327 : tRNA_Me_trans |
|
5294 : hypothetical protein |
22139 : Cyt_C5_DNA_methylase superfamily |
30454 : Brix |
38776 : BOP1NT |
|
5532 : hypothetical protein |
22200 : hypothetical protein |
30691 : AdoMet_MTases |
39022 : COG2319 |
|
6250 : reductase |
22599 : hypothetical protein |
31047 : Brix |
40044 : (MAP1) PLN03158 |
|
6803 : (RPE1) RPE |
22861 : hypothetical protein |
31415 : hypothetical protein |
40223 : PLN02347 |
|
7151 : hypothetical protein |
22985 : WD40 superfamily |
31938 : Pumilio |
40341 : WD40 superfamily |
|
7742 : KTI12 |
23106 : hypothetical protein |
32066 : Mito_carr |
42515 : DEADc |
|
8259 : hypothetical protein |
23354 : RNA-binding protein of the pumilio family |
32788 : Terminase_GpA superfamily |
261109 : NBD_sugar-kinase_HSP70_actin superfamily |
|
8997 : SrmB |
23718 : hypothetical protein |
32790 : RRM_eIF3G_like |
261462 : hypothetical protein |
|
10853 : hypothetical protein |
24038 : hypothetical protein |
32816 : DUF155 superfamily |
268048 : MFS |
|
13224 : (pabA) PABP-1234 |
24123 : fumC |
33589 : Brix superfamily |
268629 : PseudoU_synth_RluCD_like |
|
15688 : MFS |
24358 : RAP |
33718 : Methyltransf_4 superfamily |
268678 : (Tp_Myb2R3) ZUO1 |
|
17110 : rbgA |
24632 : PDCD2_C superfamily |
33871 : MPP_PP5_C |
268688 : hypothetical protein |
|
20629 : YCII superfamily |
24867 : hypothetical protein |
34348 : SAM_decarbox superfamily |
268745 : hypothetical protein |
|
20788 : Diphthine_synthase |
25277 : hypothetical protein |
34413 : RRM_ABT1_like |
268807 : (RIR1_2) PLN02437 |
|
20814 : hypothetical protein |
25412 : Nitro_FMN_reductase superfamily |
35207 : YihA_EngB |
268970 : PGDH_like_3 |
|
20879 : (Tp_CCCH21) regulator [Rayko] |
25799 : TYW3 superfamily |
35911 : COG1092 |
270145 : (LASPO) NadB |
|
21050 : Ras_like_GTPase superfamily |
26071 : (PSB_1) Ntn_hydrolase superfamily |
36053 : Pkinase |
bd1709 : (bd1709) NA |
|
21397 : Nop52 superfamily |
26678 : (TKT1) PLN02790 |
36186 : iojap_ybeB |
bd621 : (bd621) NA |
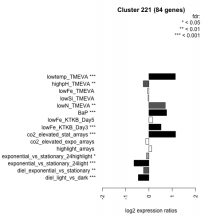
| Condition | Condition | Difference | FDR |
|---|---|---|---|
| diel_light_vs_dark | diel_light_vs_dark | -0.454 | 0.000485 |
| lowFe_KTKB_Day3 | lowFe_KTKB_Day3 | 0.520 | 0.000862 |
| lowFe_KTKB_Day5 | lowFe_KTKB_Day5 | 0.159 | 0.0966 |
| BaP | BaP | 0.761 | 0.00037 |
| exponential_vs_stationary_24highlight | exponential_vs_stationary_24highlight | -0.103 | 0.0349 |
| co2_elevated_stat_arrays | co2_elevated_stat_arrays | 1.140 | 0.000658 |
| lowtemp_TMEVA | lowtemp_TMEVA | 1.130 | 0.000735 |
| highpH_TMEVA | highpH_TMEVA | -0.241 | 0.00137 |
| co2_elevated_expo_arrays | co2_elevated_expo_arrays | -0.131 | 0.174 |
| lowFe_TMEVA | lowFe_TMEVA | -0.058 | 0.888 |
| exponential_vs_stationary_24light | exponential_vs_stationary_24light | -0.638 | 0.000581 |
| lowN_TMEVA | lowN_TMEVA | 0.702 | 0.00119 |
| diel_exponential_vs_stationary | diel_exponential_vs_stationary | -0.226 | 0.00425 |
| lowSi_TMEVA | lowSi_TMEVA | -0.034 | 1 |
| highlight_arrays | highlight_arrays | 0.117 | 0.173 |