Module 301 Residual: 0.61
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0301 | v02 | 0.61 | -8.24 |
| Rv0201c | Rv0202c Transmembrane transport protein MmpL13 |
| Rv0203 POSSIBLE EXPORTED PROTEIN | Rv0583c Lipoprotein LpqN |
| Rv0633c Possible membrane protein | Rv0951 Succinyl-CoA ligase [ADP-forming] beta chain (EC 6.2.1.5) |
| Rv2450c Putative saccharopine dehydrogenase | Rv2584c Adenine phosphoribosyltransferase (EC 2.4.2.7) |
| Rv3274c Butyryl-CoA dehydrogenase (EC 1.3.99.2) | Rv3472 |
| Rv3485c short chain dehydrogenase | Rv0547c \Oxidoreductase, short-chain dehydrogenase/reductase family\\\"" |
| Rv0952 Succinyl-CoA ligase [ADP-forming] alpha chain (EC 6.2.1.5) | Rv2352c PPE family protein |
| Rv2725c GTP-binding protein HflX | Rv3036c Possible membrane protein |
| Rv3436c Glucosamine--fructose-6-phosphate aminotransferase [isomerizing] (EC 2.6.1.16) | Rv3440c |
| Module GO Term Enrichment | Descriptions |
|---|---|
| GO:0070838 | GO:0072511 | GO:0000041 | GO:0072509 | divalent metal ion transport | divalent inorganic cation transport | transition metal ion transport | divalent inorganic cation transmembrane ... |
|
GO:0044119 | growth of symbiont in host cell |
|
GO:0005618 | cell wall |
|
GO:0005829 | cytosol |
|
GO:0005886 | plasma membrane |
|
GO:0005576 | extracellular region |
|
GO:0004775 | succinate-CoA ligase (ADP-forming) activity |
|
GO:0040007 | growth |
|
GO:0010628 | positive regulation of gene expression |
|
GO:0010629 | negative regulation of gene expression |
|
GO:0040010 | positive regulation of growth rate |
|
GO:0003999 | adenine phosphoribosyltransferase activity |
|
GO:0004085 | butyryl-CoA dehydrogenase activity |
|
GO:0005525 | GTP binding |
|
GO:0005622 | intracellular |
|
GO:0004360 | glutamine-fructose-6-phosphate transaminase (isomerizing) activity |





















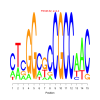
Discussion