Module 200 Residual: 0.45
| Title | Bicluster Model | Residual | Score |
|---|---|---|---|
| bicluster_0200 | v02 | 0.45 | -27.62 |
| Rv3504 FadE30 | Rv3505 Acyl-CoA dehydrogenase, short-chain specific (EC 1.3.99.2) |
| Rv3515c Long-chain-fatty-acid--CoA ligase (EC 6.2.1.3) | Rv3516 Enoyl-CoA hydratase (EC 4.2.1.17) |
| Rv3527 | Rv3570c POSSIBLE OXIDOREDUCTASE |
| Rv3573c Acyl-CoA dehydrogenase (EC 1.3.99.-) | Rv3526 Terminal oxygenase KshA |
| Rv3540c Lipid carrier protein IgrF | Rv3543c Probable acyl-CoA dehydrogenase FadE29 (EC 1.3.99.-); Acyl-CoA dehydrogenase IgrC |
| Rv3544c Probable acyl-CoA dehydrogenase FadE28 (EC 1.3.99.-); Acyl-CoA dehydrogenase IgrB | Rv3545c Putative cytochrome P450 125 (EC 1.14.-.-); Putative cytochrome P450 IgrA |
| Rv3546 Probable acetyl-CoA acetyltransferase FadA5 (EC 2.3.1.9) | Rv3568c 2,3-dihydroxybiphenyl 1,2-dioxygenase |
| Rv3574 Transcriptional regulator kstR (Rv3574), TetR family |
| Module GO Term Enrichment | Descriptions |
|---|---|
| GO:0050662 | GO:0016491 | coenzyme binding | oxidoreductase activity |
|
GO:0005886 | plasma membrane |
|
GO:0004467 | long-chain fatty acid-CoA ligase activity |
|
GO:0052572 | response to host immune response |
|
GO:0004300 | enoyl-CoA hydratase activity |
|
GO:0044117 | growth of symbiont in host |
|
GO:0070723 | response to cholesterol |
|
GO:0004085 | butyryl-CoA dehydrogenase activity |
|
GO:0009268 | response to pH |
|
GO:0009405 | pathogenesis |
|
GO:0003988 | acetyl-CoA C-acyltransferase activity |
|
GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen |
|
GO:0003677 | DNA binding |
|
GO:0042803 | protein homodimerization activity |

















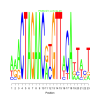
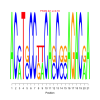
Discussion